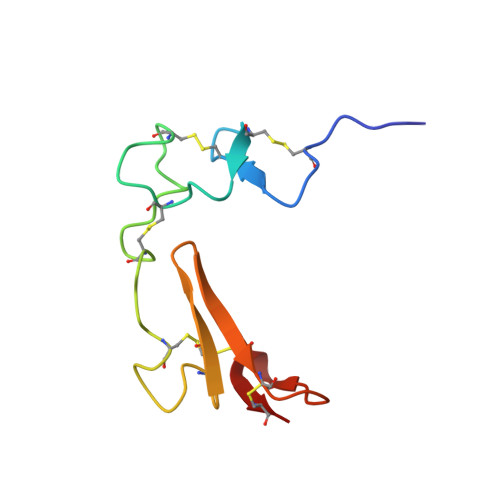
Cooperation between Fixed and Low pH-Inducible Interfaces Controls Lipoprotein Release by the LDL Receptor
Beglova, N., Jeon, H., Fisher, C., Blacklow, S.C.(2004) Mol Cell 16: 281-292
- PubMed: 15494314 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.09.038
- Primary Citation Related Structures:
1XFE - PubMed Abstract:
Low-density lipoprotein (LDL) receptors bind lipoprotein particles at the cell surface and release them in the low pH environment of the endosome. The published structure of the receptor determined at endosomal pH reveals an interdomain interface between its beta propeller and its fourth and fifth ligand binding (LA) repeats, suggesting that the receptor adopts a closed conformation at low pH to release LDL. Here, we combine lipoprotein binding and release assays with NMR spectroscopy to examine structural features of the receptor promoting release of LDL at low pH. These studies lead to a model in which the receptor uses a pH-invariant scaffold as an anchor to restrict conformational search space, combining it with flexible linkers between ligand binding repeats to interconvert between open and closed conformations. This finely tuned balance between interdomain rigidity and flexibility is likely to represent a shared structural feature in proteins of the LDL receptor family.
- Department of Pathology, Brigham and Women's Hospital and Harvard Medical School, 77 Avenue Louis Pasteur, Boston, MA 02115, USA.
Organizational Affiliation: