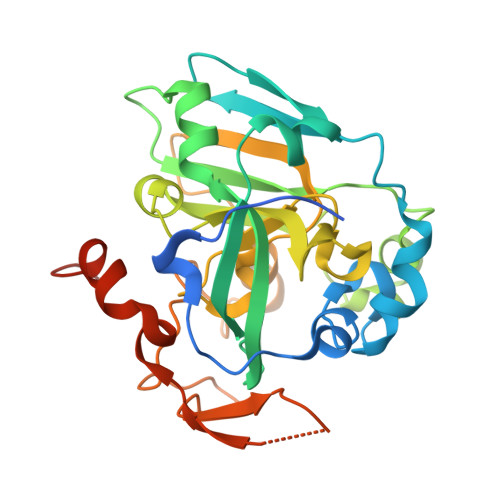
Structural and biochemical identification of a novel bacterial oxidoreductase.
Loschi, L., Brokx, S.J., Hills, T.L., Zhang, G., Bertero, M.G., Lovering, A.L., Weiner, J.H., Strynadka, N.C.(2004) J Biological Chem 279: 50391-50400
- PubMed: 15355966 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M408876200
- Primary Citation Related Structures:
1XDQ, 1XDY - PubMed Abstract:
By using a bioinformatics screen of the Escherichia coli genome for potential molybdenum-containing enzymes, we have identified a novel oxidoreductase conserved in the majority of Gram-negative bacteria. The identified operon encodes for a proposed heterodimer, YedYZ in Escherichia coli, consisting of a soluble catalytic subunit termed YedY, which is likely anchored to the membrane by a heme-containing trans-membrane subunit termed YedZ. YedY is uniquely characterized by the presence of one molybdenum molybdopterin not conjugated by an additional nucleotide, and it represents the only molybdoenzyme isolated from E. coli characterized by the presence of this cofactor form. We have further characterized the catalytic subunit YedY in both the molybdenum- and tungsten-substituted forms by using crystallographic analysis. YedY is very distinct in overall architecture from all known bacterial reductases but does show some similarity with the catalytic domain of the eukaryotic chicken liver sulfite oxidase. However, the strictly conserved residues involved in the metal coordination sphere and in the substrate binding pocket of YedY are strikingly different from that of chicken liver sulfite oxidase, suggesting a catalytic activity more in keeping with a reductase than that of a sulfite oxidase. Preliminary kinetic analysis of YedY with a variety of substrates supports our proposal that YedY and its many orthologues may represent a new type of membrane-associated bacterial reductase.
- Department of Biochemistry, University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: