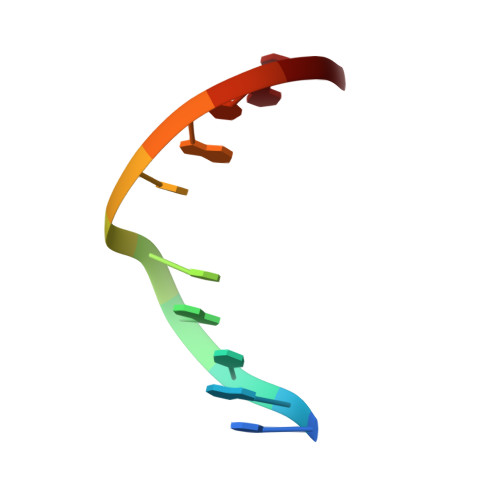
Impact of the C1 configuration of abasic sites on DNA duplex structure
de Los Santos, C., El-Khateeb, M., Rege, P., Tian, K., Johnson, F.(2004) Biochemistry 43: 15349-15357
- PubMed: 15581347 Search on PubMed
- DOI: https://doi.org/10.1021/bi048400c
- Primary Citation Related Structures:
1XCY, 1XCZ - PubMed Abstract:
Naturally occurring abasic sites in DNA exist as an equilibrium mixture of the aldehyde, the hydrated aldehyde, and the hemiacetal forms (dominant). The influence of the configuration of the C1' hydroxyl group of the hemiacetal form on duplex structure and abasic site repair has been examined using novel carbocyclic analogues. Both the alpha- and beta-forms of this novel abasic site were introduced into oligomeric DNA using the standard DMT-phosphoramidite approach in an automated solid-phase synthesizer. Solution structures of the d(CGTACXCATGC).d(GCATGAGTACG) duplex (where X is the alpha- or beta-anomer of the carbocyclic abasic site analogue) were determined by NMR spectroscopy and restrained molecular dynamics simulations. The structures were only minimally perturbed by the presence of either anomer of the abasic site. All residues adopted an anti conformation, and Watson-Crick alignments were observed on all base pairs of the duplexes. At the lesion site, the abasic residues and their partner adenines showed increased dynamic behavior but adopted intrahelical positions in the final refined structures. Incision of duplexes having the alpha- or beta-anomer of the carbocyclic abasic site by human AP endonuclease showed that the enzyme recognizes both configurations of the lesion and nicks the DNA backbone with similar efficiency. Our results challenge the suggestion that Ape1 is stereoselective and imply a plasticity at the active site of the enzyme for accommodating either anomer of the lesion.
- Department of Pharmacological Sciences, School of Medicine, State University of New York, Stony Brook, New York 11794-8651, USA. cds@pharm.sunysb.edu
Organizational Affiliation: