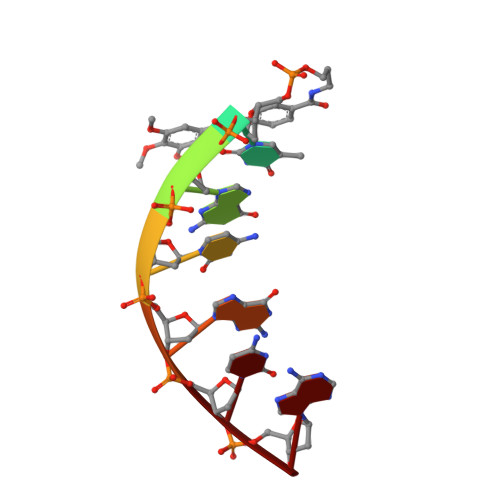
How much pi-stacking do DNA termini seek? Solution structure of a self-complementary DNA hexamer with trimethoxystilbenes capping the terminal base pairs.
Tuma, J., Paulini, R., Rojas Stutz, J.A., Richert, C.(2004) Biochemistry 43: 15680-15687
- PubMed: 15595824 Search on PubMed
- DOI: https://doi.org/10.1021/bi048205y
- Primary Citation Related Structures:
1X6W - PubMed Abstract:
The exposed terminal base pairs of DNA duplexes are nonclassical binding sites for small molecules. Instead, small molecules usually prefer intercalation or minor groove binding. Here we report the solution structure of the DNA duplex (TMS-TGCGCA)(2), where TMS denotes trimethoxystilbene carboxamides that are 5'-tethered to the DNA. The stilbenes, for which intercalation is conformationally accessible, stack on the terminal T:A base pairs of an undisturbed B-form duplex. Two conformations, differing by the orientation of the stilbene relative to the terminal base pair, are observed, indicating that the flip rate is slow for the pi-stacked aromatic ring system. The trimethoxystilbene is known to greatly increase base pairing fidelity at the terminus. Here we show that it gauges the size of the T:A base pair by embracing the 2'-methylene group of the terminal dA residue of the unmodified terminus with its methoxy "arms", but that it does not engage the entire base pair in pi-stacking. Mismatched base pairs with their altered geometry will not allow for the same embracing interaction. On the basis of the current structure, a trimethoxychrysene carboxamide is proposed as a ligand with increased pi-stacking surface and possible applications as improved fidelity-enhancing element.
- Max-Planck-Institute for Biophysical Chemistry, D-37077 Göttingen, Germany.
Organizational Affiliation: