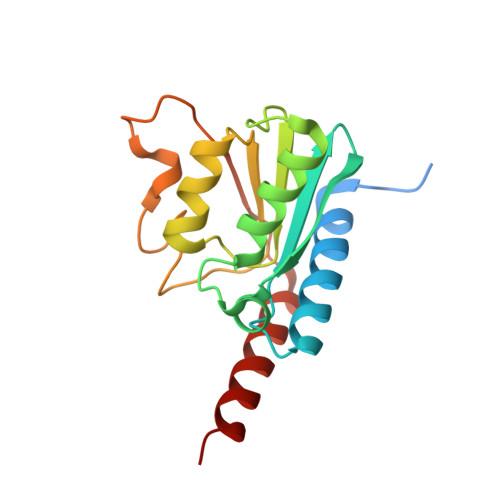
Crystal Structure of tRNA Adenosine Deaminase (TadA) from Aquifex aeolicus
Kuratani, M., Ishii, R., Bessho, Y., Fukunaga, R., Sengoku, T., Shirouzu, M., Sekine, S., Yokoyama, S.(2005) J Biological Chem 280: 16002-16008
- PubMed: 15677468 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M414541200
- Primary Citation Related Structures:
1WWR - PubMed Abstract:
The bacterial tRNA adenosine deaminase (TadA) generates inosine by deaminating the adenosine residue at the wobble position of tRNA(Arg-2). This modification is essential for the decoding system. In this study, we determined the crystal structure of Aquifex aeolicus TadA at a 1.8-A resolution. This is the first structure of a deaminase acting on tRNA. A. aeolicus TadA has an alpha/beta/alpha three-layered fold and forms a homodimer. The A. aeolicus TadA dimeric structure is completely different from the tetrameric structure of yeast CDD1, which deaminates mRNA and cytidine, but is similar to the dimeric structure of yeast cytosine deaminase. However, in the A. aeolicus TadA structure, the shapes of the C-terminal helix and the regions between the beta4 and beta5 strands are quite distinct from those of yeast cytosine deaminase and a large cavity is produced. This cavity contains many conserved amino acid residues that are likely to be involved in either catalysis or tRNA binding. We made a docking model of TadA with the tRNA anticodon stem loop.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: