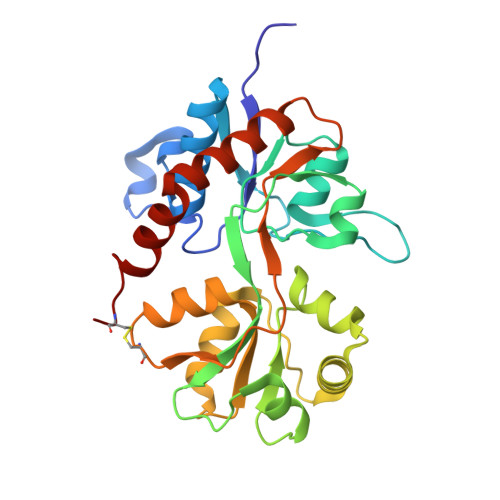
Exploring the GluR2 ligand-binding core in complex with the bicyclical AMPA analogue (S)-4-AHCP
Nielsen, B.B., Pickering, D.S., Greenwood, J.R., Brehm, L., Gajhede, M., Schousboe, A., Kastrup, J.S.(2005) FEBS J 272: 1639-1648
- PubMed: 15794751 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2005.04583.x
- Primary Citation Related Structures:
1WVJ - PubMed Abstract:
The X-ray structure of the ionotropic GluR2 ligand-binding core (GluR2-S1S2J) in complex with the bicyclical AMPA analogue (S)-2-amino-3-(3-hydroxy-7,8-dihydro-6H-cyclohepta[d]-4-isoxazolyl)propionic acid [(S)-4-AHCP] has been determined, as well as the binding pharmacology of this construct and of the full-length GluR2 receptor. (S)-4-AHCP binds with a glutamate-like binding mode and the ligand adopts two different conformations. The K(i) of (S)-4-AHCP at GluR2-S1S2J was determined to be 185 +/- 29 nM and at full-length GluR2(R)o it was 175 +/- 8 nM. (S)-4-AHCP appears to elicit partial agonism at GluR2 by inducing an intermediate degree of domain closure (17 degrees). Also, functionally (S)-4-AHCP has an efficacy of 0.38 at GluR2(Q)i, relative to (S)-glutamate. The proximity of bound (S)-4-AHCP to domain D2 prevents full D1-D2 domain closure, which is limited by steric repulsion, especially between Leu704 and the ligand.
- Biostructural Research, Department of Medicinal Chemistry, Danish University of Pharmaceutical Sciences, Copenhagen, Denmark.
Organizational Affiliation: