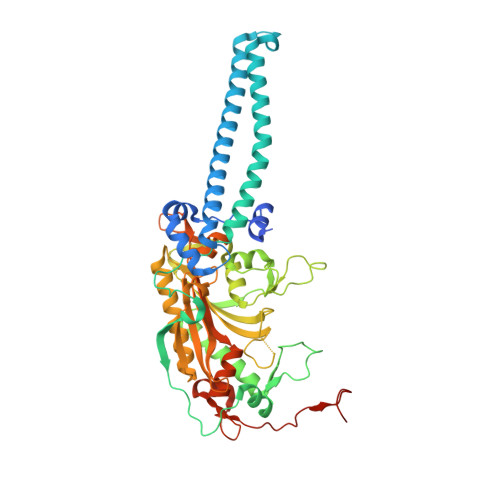
Dual-mode recognition of noncanonical tRNAs(Ser) by seryl-tRNA synthetase in mammalian mitochondria
Chimnaronk, S., Jeppesen, M.G., Suzuki, T., Nyborg, J., Watanabe, K.(2005) EMBO J 24: 3369-3379
- PubMed: 16163389 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600811
- Primary Citation Related Structures:
1WLE - PubMed Abstract:
The secondary structures of metazoan mitochondrial (mt) tRNAs(Ser) deviate markedly from the paradigm of the canonical cloverleaf structure; particularly, tRNA(Ser)(GCU) corresponding to the AGY codon (Y=U and C) is highly truncated and intrinsically missing the entire dihydrouridine arm. None of the mt serine isoacceptors possesses the elongated variable arm, which is the universal landmark for recognition by seryl-tRNA synthetase (SerRS). Here, we report the crystal structure of mammalian mt SerRS from Bos taurus in complex with seryl adenylate at an atomic resolution of 1.65 A. Coupling structural information with a tRNA-docking model and the mutagenesis studies, we have unraveled the key elements that establish tRNA binding specificity, differ from all other known bacterial and eukaryotic systems, are the characteristic extensions in both extremities, as well as a few basic residues residing in the amino-terminal helical arm of mt SerRS. Our data further uncover an unprecedented mechanism of a dual-mode recognition employed to discriminate two distinct 'bizarre' mt tRNAs(Ser) by alternative combination of interaction sites.
- Department of Integrated Biosciences, Graduate School of Frontier Sciences, University of Tokyo, Kashiwanoha, Kashiwa, Chiba, Japan.
Organizational Affiliation: