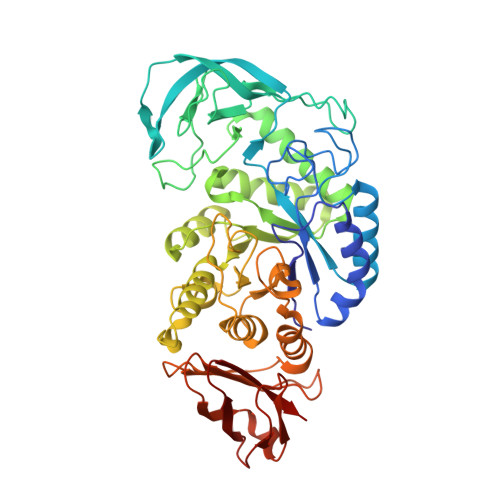
Structure of a Bacillus Halmapalus Family 13 Alpha-Amylase, Bha, in Complex with an Acarbose-Derived Nonasaccharide at 2.1 A Resolution
Davies, G.J., Brzozowski, A.M., Dauter, Z., Rasmussen, M.D., Borchert, T.V., Wilson, K.S.(2005) Acta Crystallogr D Biol Crystallogr 61: 190
- PubMed: 15681870 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904027118
- Primary Citation Related Structures:
1W9X - PubMed Abstract:
The enzymatic digestion of starch by alpha-amylases is one of the key biotechnological reactions of recent times. In the search for industrial biocatalysts, the family GH13 alpha-amylase BHA from Bacillus halmapalus has been cloned and expressed. The three-dimensional structure at 2.1 A resolution has been determined in complex with the (pseudo)tetrasaccharide inhibitor acarbose. Acarbose is found bound as a nonasaccharide transglycosylation product spanning the -6 to +3 subsites. Careful inspection of electron density suggests that the bound ligand could not have been formed through successive transglycosylations of acarbose and must also have featured maltose or maltooligosaccharides as an acceptor.
- Department of Chemistry, University of York, England. davies@ysbl.york.ac.uk
Organizational Affiliation: