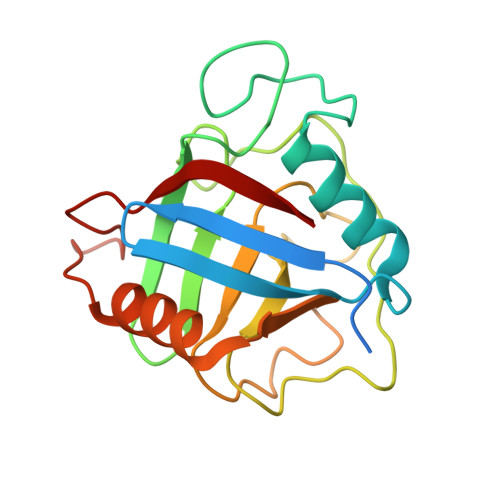
X-Ray Structure of Peptidyl-Prolyl Cis-Trans Isomerase a from Mycobacterium Tuberculosis
Henriksson, L.M., Johansson, P., Unge, T., Mowbray, S.L.(2004) Eur J Biochem 271: 4107
- PubMed: 15479239 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.2004.04348.x
- Primary Citation Related Structures:
1W74 - PubMed Abstract:
Peptidyl-prolyl cis-trans isomerases (EC 5.2.1.8) catalyse the interconversion of cis and trans peptide bonds and are therefore considered to be important for protein folding. They are also thought to participate in processes such as signalling, cell surface recognition, chaperoning and heat-shock response. Here we report the soluble expression of recombinant Mycobacterium tuberculosis peptidyl-prolyl cis-trans isomerase PpiA in Escherichia coli, together with an investigation of its structure and biochemical properties. The protein was shown to be active in a spectrophotometric assay, with an estimated kcat/Km of 2.0 x 10(6) m(-1).s(-1). The X-ray structure of PpiA was solved by molecular replacement, and refined to a resolution of 2.6 A with R and Rfree values of 21.3% and 22.9%, respectively. Comparisons to known structures show that the PpiA represents a slight variation on the peptidyl-prolyl cis-trans isomerase fold, previously not represented in the Protein Data Bank. Inspection of the active site suggests that specificity for substrates and cyclosporin A will be similar to that found for most other enzymes of this structural family. Comparison to the sequence of the second M. tuberculosis enzyme, PpiB, suggests that binding of peptide substrates as well as cyclosporin A may differ in that case.
- Department of Cell and Molecular Biology, Uppsala University, Sweden.
Organizational Affiliation: