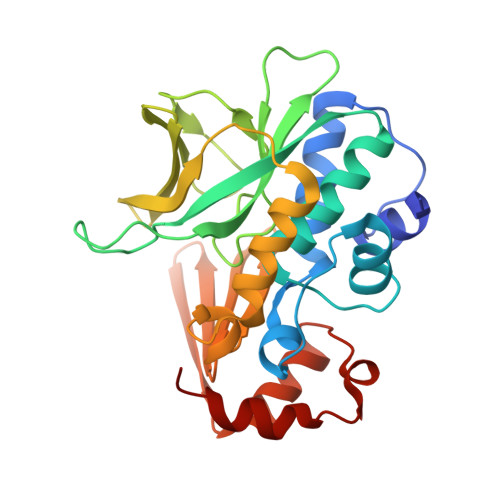
Binding of the Anti-Tubercular Drug Isoniazid to the Arylamine N-Acetyltransferase Protein from Mycobacterium Smegmatis
Sandy, J., Holton, S., Fullham, E., Sim, E., Noble, M.E.M.(2005) Protein Sci 14: 775
- PubMed: 15722451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.041163505
- Primary Citation Related Structures:
1W6F - PubMed Abstract:
Isoniazid is a frontline drug used in the treatment of tuberculosis (TB). Isoniazid is a prodrug, requiring activation in the mycobacterial cell by the catalase/peroxidase activity of the katG gene product. TB kills two million people every year and the situation is getting worse due to the increase in prevalence of HIV/AIDS and emergence of multidrug-resistant strains of TB. Arylamine N-acetyltransferase (NAT) is a drug-metabolizing enzyme (E.C. 2.1.3.5). NAT can acetylate isoniazid, transferring an acetyl group from acetyl coenzyme A onto the terminal nitrogen of the drug, which in its N-acetylated form is therapeutically inactive. The bacterium responsible for TB, Mycobacterium tuberculosis, contains and expresses the gene encoding the NAT protein. Isoniazid binds to the NAT protein from Salmonella typhimurium and we report here the mode of binding of isoniazid in the NAT enzyme from Mycobacterium smegmatis, closely related to the M. tuberculosis and S. typhimurium NAT enzymes. The mode of binding of isoniazid to M. smegmatis NAT has been determined using data collected from two distinct crystal forms. We can say with confidence that the observed mode of binding of isoniazid is not an artifact of the crystallization conditions used. The NAT enzyme is active in mycobacterial cells and we propose that isoniazid binds to the NAT enzyme in these cells. NAT activity in M. tuberculosis is likely therefore to modulate the degree of activation of isoniazid by other enzymes within the mycobacterial cell. The structure of NAT with isoniazid bound will facilitate rational drug design for anti-tubercular therapy.
- Department of Pharmacology, Oxford University, Mansfield Road, Oxford, OX1 3QT, United Kingdom. james.sandy@pharm.ox.ac.uk
Organizational Affiliation: