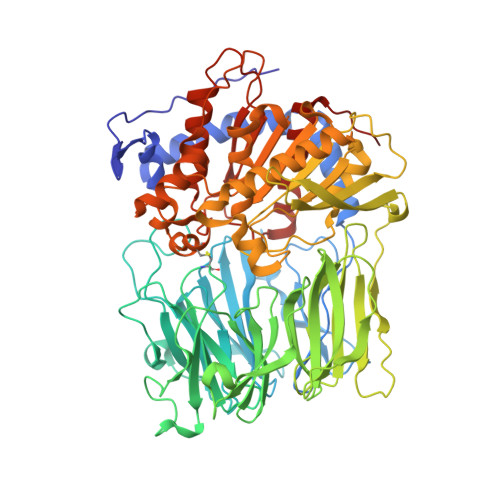
Concerted Structural Changes in the Peptidase and the Propeller Domains of Prolyl Oligopeptidase are Required for Substrate Binding
Szeltner, Z., Rea, D., Juhasz, T., Renner, V., Fulop, V., Polgar, L.(2004) J Mol Biology 340: 627
- PubMed: 15210359 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.05.011
- Primary Citation Related Structures:
1VZ2, 1VZ3 - PubMed Abstract:
Prolyl oligopeptidase contains a peptidase domain and its catalytic triad is covered by the central tunnel of a seven-bladed beta-propeller. This domain makes the enzyme an oligopeptidase by excluding large structured peptides from the active site. The apparently rigid crystal structure does not explain how the substrate can approach the catalytic groups. Two possibilities of substrate access were investigated: either blades 1 and 7 of the propeller domain move apart, or the peptidase and/or propeller domains move to create an entry site at the domain interface. Engineering disulfide bridges to the expected oscillating structures prevented such movements, which destroyed the catalytic activity and precluded substrate binding. This indicated that concerted movements of the propeller and the peptidase domains are essential for the enzyme action.
- Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences, H-1518 Budapest 112, P.O. Box 7, Hungary.
Organizational Affiliation: