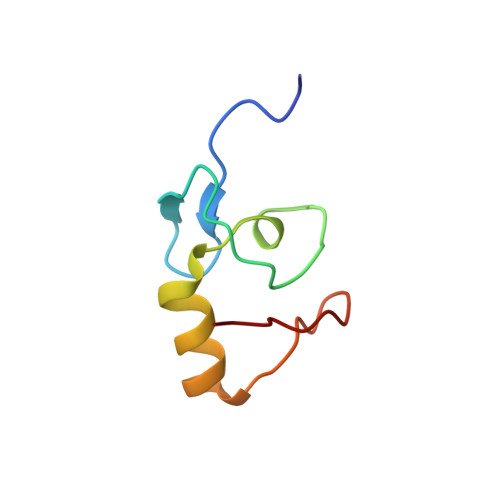
Solution Structure of the Kaposi'S Sarcoma-Associated Herpesvirus K3 N-Terminal Domain Reveals a Novel E2-Binding C4Hc3-Type Ring Domain
Dodd, R.B., Allen, M.D., Brown, S.E., Sanderson, C.M., Duncan, L.M., Lehner, P.J., Bycroft, M., Read, R.J.(2004) J Biological Chem 279: 53840
- PubMed: 15465811 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409662200
- Primary Citation Related Structures:
1VYX - PubMed Abstract:
RING domains are found in a large number of eukaryotic proteins. Most function as E3 ubiquitin-protein ligases, catalyzing the terminal step in the ubiquitination process. Structurally, these domains have been characterized as binding two zinc ions in a stable cross-brace motif. The tumorigenic human gamma-herpesvirus Kaposi's sarcoma-associated herpesvirus encodes a ubiquitin-protein ligase termed K3, which functions as an immune evasion molecule by ubiquitinating major histocompatibility complex class I. K3 possesses at its N terminus a domain related to cellular RING domains but with an altered zinc ligand arrangement. This domain was initially characterized as a plant homeodomain, a structure not previously known to function as an E3. Here, it is conclusively demonstrated that the K3 N-terminal domain is a variant member of the RING domain family and not a plant homeodomain. The domain is found to interact with the cellular ubiquitin-conjugating enzymes UbcH5A to -C and UbcH13, which dock to the equivalent surface as on classical cellular RING domains. Interaction with UbcH13 suggests a possible role for K3 in catalyzing Lys(63)-linked ubiquitination.
- Cambridge Institute for Medical Research, Wellcome Trust/MRC Building, Addenbrooke's Hospital, Hills Road, Cambridge CB2 2XY, UK.
Organizational Affiliation: