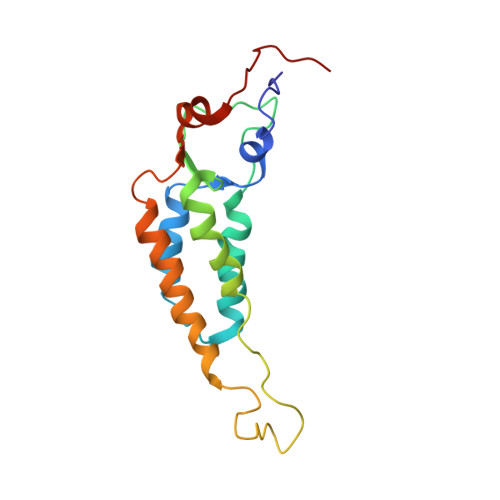
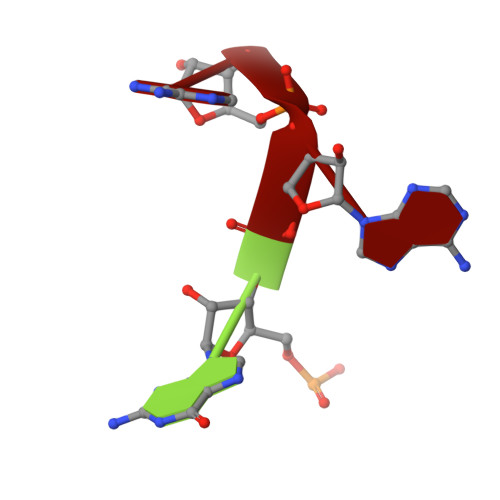
Structure of the U2 strain of tobacco mosaic virus refined at 3.5 A resolution using X-ray fiber diffraction.
Pattanayek, R., Stubbs, G.(1992) J Mol Biology 228: 516-528
- PubMed: 1453461 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90839-c
- Primary Citation Related Structures:
1VTM - PubMed Abstract:
The structure of the U2 strain of tobacco mosaic virus (TMV) has been determined by fiber diffraction methods at 3.5 A resolution, and refined by a combination of restrained least-squares and molecular dynamics methods to an R-factor of 0.096. The structure is extremely similar to that of the common strain of TMV, with the largest differences being in the protein loop that makes up the inner surface of the virus, and in the C-terminal region on the outer surface. Differences in the inner loop can be correlated with differences in the properties of the two viruses.
- Department of Molecular Biology Vanderbilt University, Nashville, TN 37235.
Organizational Affiliation: