Interactions of Quinoxaline Antibiotic and DNA: The Molecular Structure of a Triostin A-D(Gcgtacgc) Complex.
Wang, A.H., Ughetto, G., Quigley, G.J., Rich, A.(1986) J Biomol Struct Dyn 4: 319
- PubMed: 3271447 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.1986.10506353
- Primary Citation Related Structures:
1VS2 - PubMed Abstract:
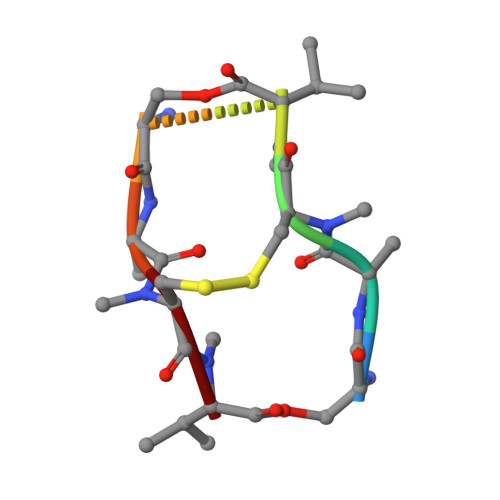
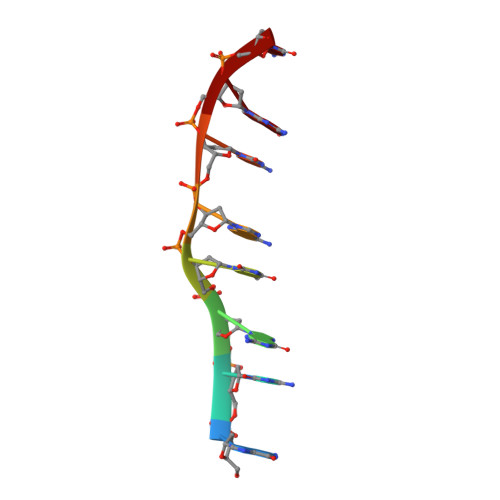
The crystal structure of a DNA octamer d(GCGTACGC) complexed to an antitumor antibiotic, triostin A, has been solved and refined to 2.2 A resolution by x-ray diffraction analysis. The antibiotic molecule acts as a true bis intercalator surrounding the d(CpG) sequence at either end of the unwound right-handed DNA double helix. As previously observed in the structure of triostin A-d(CGTACG) complex (A.H.-J. Wang, et. al., Science, 225, 1115-1121 (1984)), the alanine amino acid residues of the drug molecule form sequence-specific hydrogen bonds to guanines in the minor groove. The two central A.T base pairs are in Hoogsteen configuration with adenine in the syn conformation. In addition, the two terminal G.C base pairs flanking the quinoxaline rings are also held together by Hoogsteen base pairing. This is the first observation in an oligonucleotide of. Hoogsteen G.C base pairs where the cytosine is protonated. The principal functional components of a bis-intercalative compound are discussed.
- Department of Biology, Massachusetts Institute of Technology, Cambridge 02139.
Organizational Affiliation: