Identification of 1,5-Naphthyridine Derivatives as a Novel Series of Potent and Selective TGF-beta Type I Receptor Inhibitors.
Gellibert, F., Woolven, J., Fouchet, M.-H., Mathews, N., Goodland, H., Lovegrove, V., Laroze, A., Nguyen, V.-L., Sautet, S., Wang, R., Janson, C., Smith, W., Krysa, G., Boullay, V., De Gouville, A.-C., Huet, S., Hartley, D.(2004) J Med Chem 47: 4494-4506
- PubMed: 15317461 Search on PubMed
- DOI: https://doi.org/10.1021/jm0400247
- Primary Citation Related Structures:
1VJY - PubMed Abstract:
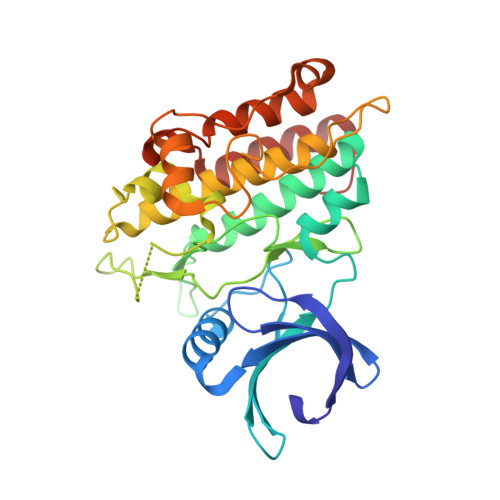
Optimization of the screening hit 1 led to the identification of novel 1,5-naphthyridine aminothiazole and pyrazole derivatives, which are potent and selective inhibitors of the transforming growth factor-beta type I receptor, ALK5. Compounds 15 and 19, which inhibited ALK5 autophosphorylation with IC50 = 6 and 4 nM, respectively, showed potent activities in both binding and cellular assays and exhibited selectivity over p38 mitogen-activated protein kinase. The X-ray crystal structure of 19 in complex with human ALK5 is described, confirming the binding mode proposed from docking studies.
- Department of Medicinal Chemistry, GlaxoSmithKline, 25-27 Avenue du Québec, 91951 Les Ulis, France. fjg23217@gsk.com
Organizational Affiliation: