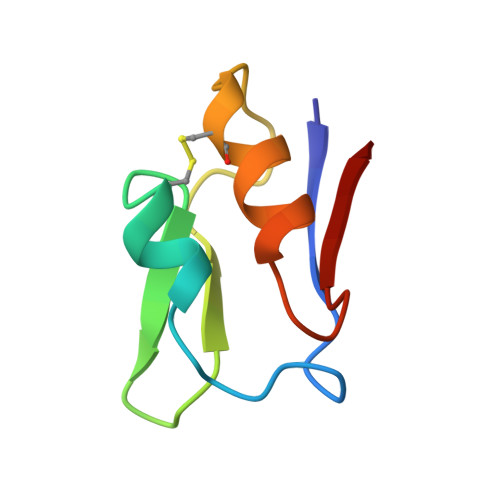
Small structural changes account for the high thermostability of 1[4Fe-4S] ferredoxin from the hyperthermophilic bacterium Thermotoga maritima.
Macedo-Ribeiro, S., Darimont, B., Sterner, R., Huber, R.(1996) Structure 4: 1291-1301
- PubMed: 8939753 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(96)00137-2
- Primary Citation Related Structures:
1VJW - PubMed Abstract:
The characterization of the structural features that account for the high thermostability of some proteins is of great scientific and biotechnological interest. Proteins from hyperthermophilic organisms with optimum growth temperatures of 80 degrees C and higher generally show high intrinsic stabilities. The comparison of high resolution X-ray structures of these proteins with their counterparts from mesophilic organisms has therefore helped to identify potentially stabilizing forces in a number of cases. Small monomeric proteins which comprise only a single domain, such as ferredoxins, are especially suitable for such comparisons since the search for determinants of protein stability is considerably simplified. The 1.75 A crystal structure of the extremely thermostable 1[4Fe-4S] ferredoxin from Thermotoga maritima (FdTm) was determined and compared with other monocluster-containing ferredoxins with different degrees of thermostability. A comparison of the three-dimensional structure of FdTm with that of ferredoxins from mesophilic organisms suggests that the very high thermostability of FdTm is unexpectedly achieved without large changes of the overall protein structure. Instead, an increased number of potentially stabilizing features is observed in FdTm, compared with mesophilic ferredoxins. These include stabilization of alpha helices, replacement of residues in strained conformation by glycines, strong docking of the N-terminal methionine and an overall increase in the number of hydrogen bonds. Most of these features stabilize several secondary structure elements and improve the overall rigidity of the polypeptide backbone. The decreased flexibility will certainly play a relevant role in shielding the iron-sulfur cluster against physiologically high temperatures and further improve the functional integrity of FdTm.
- Max-Planck Institut für Biochemie, D-82152 Martinsried, Germany. ribeiro@biochem.mpg.de
Organizational Affiliation: