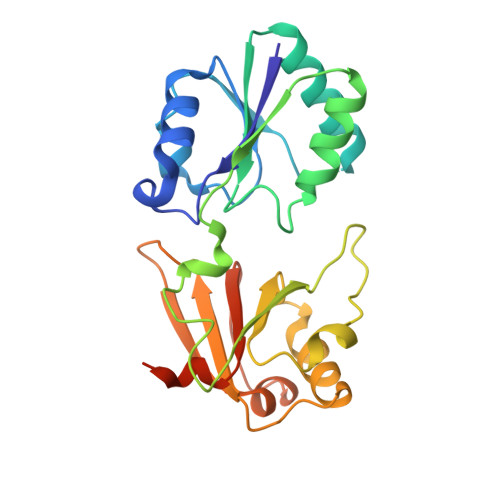
Structure of a closed-form uroporphyrinogen-III C-methyltransferase from Thermus thermophilus.
Rehse, P.H., Kitao, T., Tahirov, T.H.(2005) Acta Crystallogr D Biol Crystallogr 61: 913-919
- PubMed: 15983414 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905008838
- Primary Citation Related Structures:
1V9A, 1VA0 - PubMed Abstract:
Uroporphyrinogen-III C-methyltransferase from Thermus thermophilus is a multifunctional protein responsible for two of the eight S-adenosyl-methionine-dependent methylations of the corrin ring during vitamin B(12) synthesis. The structure of this protein has been solved to 2.0 A resolution in both the apo and cofactor-bound form. The monomer consists of two domains, A and B, each consisting of a five-stranded beta-sheet and two or three alpha-helices, with the cofactor bound at the interface. The biological unit is the dimer found in the asymmetric unit. This dimer is related by a non-crystallographic twofold such that two B domains combine to form a long ten-stranded beta-sheet. When compared with solved related structures, this structure shows clear differences in the region involved in cofactor and substrate binding, affirming the role of several previously implicated residues and questioning others. The solved related structures are characterized by an exposed active site. The T. thermophilus structure has this site restricted by the interaction of a flexible loop structure with a highly conserved residue, suggesting a mechanistic role. This structure represents the ;closed' form of the protein.
- APCG, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: