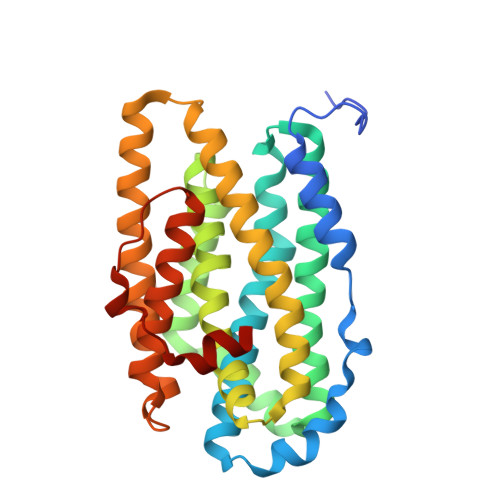
Crystal Structure of the Biologically Active Form of Class Ib Ribonucleotide Reductase Small Subunit from Mycobacterium Tuberculosis
Uppsten, M., Davis, J., Rubin, H., Uhlin, U.(2004) FEBS Lett 569: 117
- PubMed: 15225619 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2004.05.059
- Primary Citation Related Structures:
1UZR - PubMed Abstract:
Two nrdF genes of Mycobacterium tuberculosis code for different R2 subunits of the class Ib ribonucleotide reductase (RNR). The proteins are denoted R2F-1 and R2F-2 having 71% sequence identity. The R2F-2 subunit forms the biologically active RNR complex with the catalytic R1E-subunit. We present the structure of the reduced R2F-2 subunit to 2.2 A resolution. Comparison of the R2F-2 structure with a model of R2F-1 suggests that the important differences are located at the C-terminus. We found that within class Ib, the E-helix close to the iron diiron centre has two preferred conformations, which cannot be explained by the redox-state of the diiron centre. In the R2F-2 structure, we also could see a mobility of alphaE in between the two conformations.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala Biomedical Center, P.O. Box 590, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: