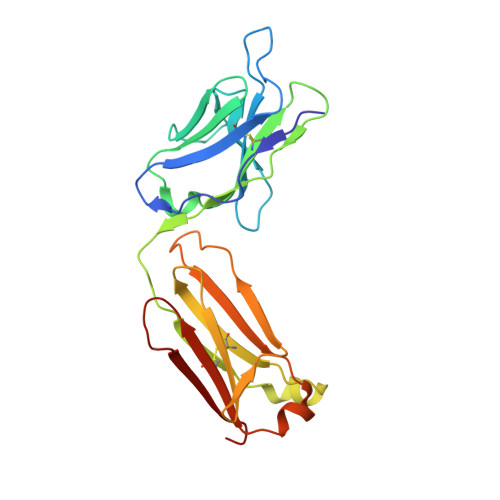
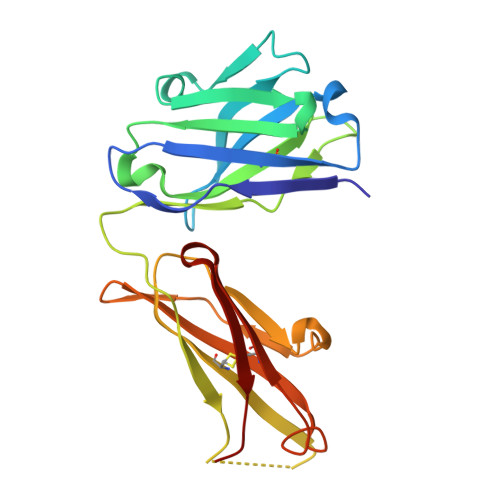
Structure of an Anti-Lewis X Fab Fragment in Complex with its Lewis X Antigen
Van Roon, A.M.M., Pannu, N.S., De Vrind, J.P.M., Van Der Marel, G.A., Van Boom, J.H., Hokke, C.H., Deelder, A.M., Abrahams, J.P.(2004) Structure 12: 1227
- PubMed: 15242599 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.05.008
- Primary Citation Related Structures:
1UZ6, 1UZ8 - PubMed Abstract:
The Lewis X trisaccharide is pivotal in mediating specific cell-cell interactions. Monoclonal antibody 291-2G3-A, which was generated from mice infected with schistosomes, has been shown to recognize the Lewis X trisaccharide. Here we describe the structure of the Fab fragment of 291-2G3-A, with Lewis X, to 1.8 A resolution. The crystallographic analysis revealed that the antigen binding site is a rather shallow binding pocket, and residues from all six complementary determining regions of the antibody contact all sugar residues. The high specificity of the binding pocket does not result in high affinity; the K(D) determined by isothermal calorimetry is 11 microM. However, this affinity is in the same range as for other sugar-antibody complexes. The detailed understanding of the antibody-Lewis X interaction revealed by the crystal structure may be helpful in the design of better diagnostic tools for schistosomiasis and for studying Lewis X-mediated cell-cell interactions by antibody interference.
- Department of Biophysical Structural Chemistry, Leiden University, P.O. Box 9502, 2300 RA Leiden, The Netherlands.
Organizational Affiliation: