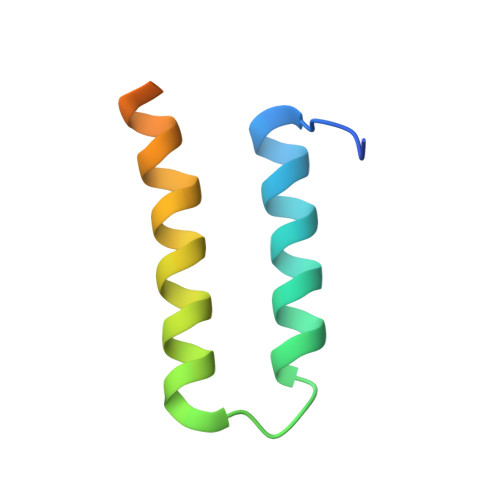
Crystal structure of the human centromere protein B (CENP-B) dimerization domain at 1.65-A resolution
Tawaramoto, M.S., Park, S.-Y., Tanaka, Y., Nureki, O., Kurumizaka, H., Yokoyama, S.(2003) J Biological Chem 278: 51454-51461
- PubMed: 14522975 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M310388200
- Primary Citation Related Structures:
1UFI - PubMed Abstract:
The human centromere protein B (CENP-B), a centromeric heterochromatin component, forms a homodimer that specifically binds to a distinct DNA sequence (the CENP-B box), which appears within every other alpha-satellite repeat. Previously, we determined the structure of the human CENP-B DNA-binding domain, CENP-B-(1-129), complexed with the CENP-B box DNA. In the present study, we determined the crystal structure of its dimerization domain (CENP-B-(540-599)), another functional domain of CENP-B, at 1.65-A resolution. CENP-B-(540-599) contains two alpha-helices, which are folded into an antiparallel configuration. The CENP-B-(540-599) dimer formed a symmetrical, antiparallel, four-helix bundle structure with a large hydrophobic patch in which 23 residues of one monomer form van der Waals contacts with the other monomer. In the CENP-B-(540-599) dimer, the N-terminal ends of CENP-B-(540-599) are oriented on opposite sides of the dimer. This CENP-B dimer configuration may be suitable for capturing two distant CENP-B boxes during centromeric heterochromatin formation.
- RIKEN Genomic Sciences Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: