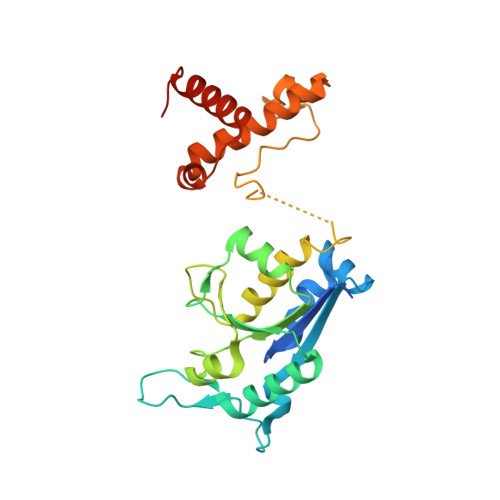
Crystal structure of tRNA(m(1)G37)methyltransferase: insights into tRNA recognition
Ahn, H.J., Kim, H.-W., Yoon, H.-J., Lee, B.I., Suh, S.W., Yang, J.K.(2003) EMBO J 22: 2593-2603
- PubMed: 12773376 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg269
- Primary Citation Related Structures:
1UAJ, 1UAK, 1UAL, 1UAM - PubMed Abstract:
tRNA(m(1)G37)methyltransferase (TrmD) catalyzes the transfer of a methyl group from S-adenosyl-L- methionine (AdoMet) to G(37) within a subset of bacterial tRNA species, which have a G residue at the 36th position. The modified guanosine is adjacent to and 3' of the anticodon and is essential for the maintenance of the correct reading frame during translation. Here we report four crystal structures of TrmD from Haemophilus influenzae, as binary complexes with either AdoMet or S-adenosyl-L-homocysteine (AdoHcy), as a ternary complex with AdoHcy and phosphate, and as an apo form. This first structure of TrmD indicates that it functions as a dimer. It also suggests the binding mode of G(36)G(37) in the active site of TrmD and the catalytic mechanism. The N-terminal domain has a trefoil knot, in which AdoMet or AdoHcy is bound in a novel, bent conformation. The C-terminal domain shows structural similarity to trp repressor. We propose a plausible model for the TrmD(2)-tRNA(2) complex, which provides insights into recognition of the general tRNA structure by TrmD.
- Structural Proteomics Laboratory, School of Chemistry and Molecular Engineering, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: