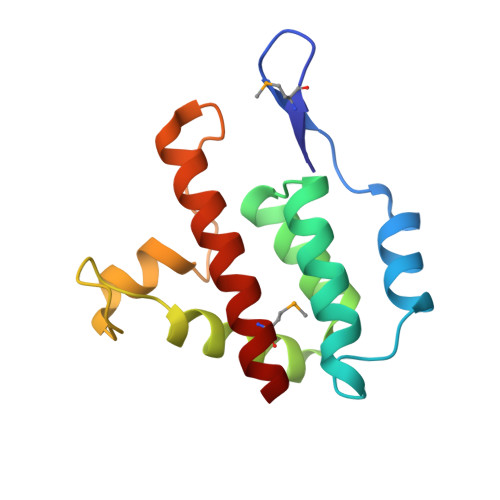
High-resolution structure of a retroviral capsid hexameric amino-terminal domain.
Mortuza, G.B., Haire, L.F., Stevens, A., Smerdon, S.J., Stoye, J.P., Taylor, I.A.(2004) Nature 431: 481-485
- PubMed: 15386017 Search on PubMed
- DOI: https://doi.org/10.1038/nature02915
- Primary Citation Related Structures:
1U7K - PubMed Abstract:
Retroviruses are the aetiological agents of a range of human diseases including AIDS and T-cell leukaemias. They follow complex life cycles, which are still only partly understood at the molecular level. Maturation of newly formed retroviral particles is an essential step in production of infectious virions, and requires proteolytic cleavage of Gag polyproteins in the immature particle to form the matrix, capsid and nucleocapsid proteins present in the mature virion. Capsid proteins associate to form a dense viral core that may be spherical, cylindrical or conical depending on the genus of the virus. Nonetheless, these assemblies all appear to be composed of a lattice formed from hexagonal rings, each containing six capsid monomers. Here, we describe the X-ray structure of an individual hexagonal assembly from N-tropic murine leukaemia virus (N-MLV). The interface between capsid monomers is generally polar, consistent with weak interactions within the hexamer. Similar architectures are probably crucial for the regulation of capsid assembly and disassembly in all retroviruses. Together, these observations provide new insights into retroviral uncoating and how cellular restriction factors may interfere with viral replication.
- Division of Protein Structure, National Institute for Medical Research, The Ridgeway, Mill Hill, London NW7 1AA, UK.
Organizational Affiliation: