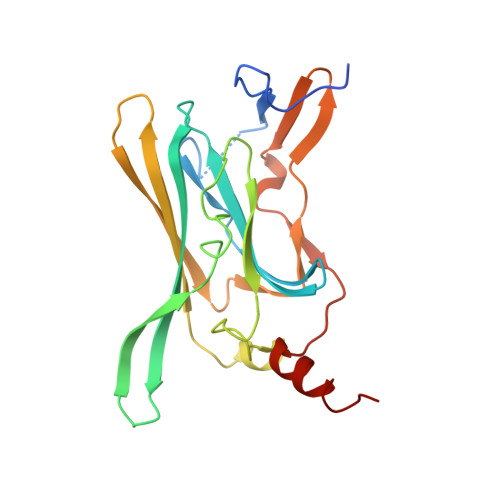
Cytoplasmic domain structures of Kir2.1 and Kir3.1 show sites for modulating gating and rectification
Pegan, S., Arrabit, C., Zhou, W., Kwiatkowski, W., Collins, A., Slesinger, P.A., Choe, S.(2005) Nat Neurosci 8: 279-287
- PubMed: 15723059 Search on PubMed
- DOI: https://doi.org/10.1038/nn1411
- Primary Citation Related Structures:
1U4E, 1U4F - PubMed Abstract:
N- and C-terminal cytoplasmic domains of inwardly rectifying K (Kir) channels control the ion-permeation pathway through diverse interactions with small molecules and protein ligands in the cytoplasm. Two new crystal structures of the cytoplasmic domains of Kir2.1 (Kir2.1(L)) and the G protein-sensitive Kir3.1 (Kir3.1(S)) channels in the absence of PIP(2) show the cytoplasmic ion-permeation pathways occluded by four cytoplasmic loops that form a girdle around the central pore (G-loop). Significant flexibility of the pore-facing G-loop of Kir2.1(L) and Kir3.1(S) suggests a possible role as a diffusion barrier between cytoplasmic and transmembrane pores. Consistent with this, mutations of the G-loop disrupted gating or inward rectification. Structural comparison shows a di-aspartate cluster on the distal end of the cytoplasmic pore of Kir2.1(L) that is important for modulating inward rectification. Taken together, these results suggest the cytoplasmic domains of Kir channels undergo structural changes to modulate gating and inward rectification.
- Structural Biology, The Salk Institute, La Jolla, California 92037, USA.
Organizational Affiliation: