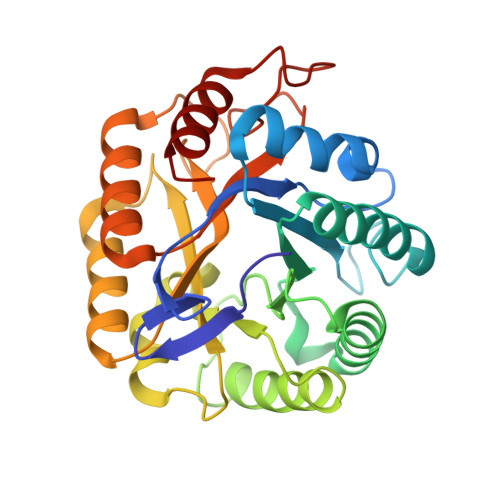
Structure of a Full Length Psychrophilic Cellulase from Pseudoalteromonas haloplanktis revealed by X-ray Diffraction and Small Angle X-ray Scattering
Violot, S., Aghajari, N., Czjzek, M., Feller, G., Sonan, G.K., Gouet, P., Gerday, C., Haser, R., Receveur-Brechot, V.(2005) J Mol Biology 348: 1211-1224
- PubMed: 15854656 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.03.026
- Primary Citation Related Structures:
1TVN, 1TVP - PubMed Abstract:
Pseudoalteromonas haloplanktis is a psychrophilic Gram-negative bacterium isolated in Antarctica, that lives on organic remains of algae. This bacterium converts the cellulose, highly constitutive of algae, into an immediate nutritive form by biodegrading this biopolymer. To understand the mechanisms of cold adaptation of its enzymatic components, we studied the structural properties of an endoglucanase, Cel5G, by complementary methods, X-ray crystallography and small angle X-ray scattering. Using X-ray crystallography, we determined the structure of the catalytic core module of this family 5 endoglucanase, at 1.4A resolution in its native form and at 1.6A in the cellobiose-bound form. The catalytic module of Cel5G presents the (beta/alpha)(8)-barrel structure typical of clan GH-A of glycoside hydrolase families. The structural comparison of the catalytic core of Cel5G with the mesophilic catalytic core of Cel5A from Erwinia chrysanthemi revealed modifications at the atomic level leading to higher flexibility and thermolability, which might account for the higher activity of Cel5G at low temperatures. Using small angle X-ray scattering we further explored the structure at the entire enzyme level. We analyzed the dimensions, shape, and conformation of Cel5G full length in solution and especially of the linker between the catalytic module and the cellulose-binding module. The results showed that the linker is unstructured, and unusually long and flexible, a peculiarity that distinguishes it from its mesophilic counterpart. Loops formed at the base by disulfide bridges presumably add constraints to stabilize the most extended conformations. These results suggest that the linker plays a major role in cold adaptation of this psychrophilic enzyme, allowing steric optimization of substrate accessibility.
- Laboratoire de BioCristallographie, Institut de Biologie et Chimie des Protéines, CNRS et Université Claude Bernard Lyon 1, UMR 5086, IFR 128 Biosciences Lyon-Gerland, 7 Passage du Vercors, F-69367 Lyon Cedex 07, France.
Organizational Affiliation: