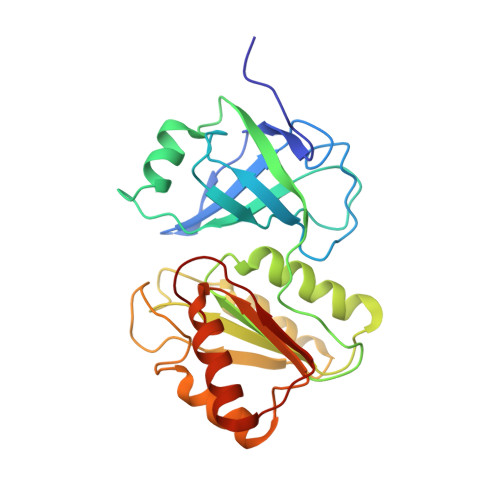
NMR Structure of the Flavin Domain from Soluble Methane Monooxygenase Reductase from Methylococcus capsulatus (Bath)
Chatwood, L.L., Gross, J.D., Wagner, G., Lippard, S.J.(2004) Biochemistry 43: 11983-11991
- PubMed: 15379538 Search on PubMed
- DOI: https://doi.org/10.1021/bi049066n
- Primary Citation Related Structures:
1TVC - PubMed Abstract:
Soluble methane monooxygenase (sMMO) catalyzes the hydroxylation of methane by dioxygen to methanol, the first step in carbon assimilation by methanotrophs. This multicomponent system transfers electrons from NADH through a reductase component to the non-heme diiron center in the hydroxylase where O(2) is activated. The reductase component comprises three distinct domains, a [2Fe-2S] ferredoxin domain along with FAD- and NADH-binding domains. We report the solution structure of the reduced 27.6 kDa FAD- and NADH-binding domains (MMOR-FAD) of the reductase from Methylococcus capsulatus (Bath). The FAD-binding domain consists of a six-stranded antiparallel beta-barrel and one alpha-helix, with the first 10 N-terminal residues unstructured. In the interface between the two domains, the FAD cofactor is tightly bound in an unprecedented extended conformation. The NADH-binding domain consists of a five-stranded parallel beta-sheet with four alpha-helices packing closely around this sheet. MMOR-FAD is structurally homologous to other FAD-containing oxidoreductases, and we expect similar structures for the FAD/NADH-binding domains of reductases that occur in other multicomponent monooxygenases.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139-4307, USA.
Organizational Affiliation: