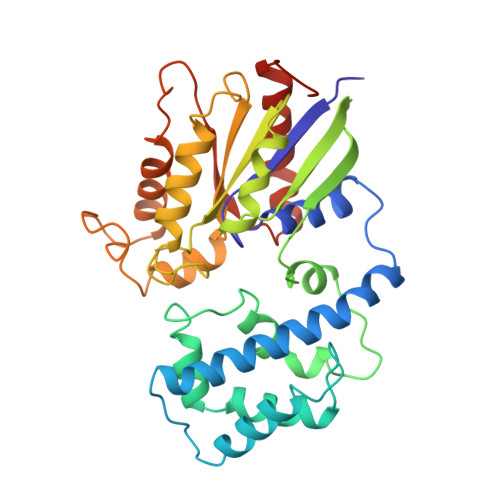
The 2.2 A crystal structure of transducin-alpha complexed with GTP gamma S.
Noel, J.P., Hamm, H.E., Sigler, P.B.(1993) Nature 366: 654-663
- PubMed: 8259210 Search on PubMed
- DOI: https://doi.org/10.1038/366654a0
- Primary Citation Related Structures:
1TND - PubMed Abstract:
The 2.2 A crystal structure of activated rod transducin, Gt alpha.GTP gamma S, shows the bound GTP gamma S molecule occluded deep in a cleft between a domain structurally homologous to small GTPases and a helical domain unique to heterotrimeric G proteins. The structure, when combined with biochemical and genetic studies, suggests: how an activated receptor might open this cleft to allow nucleotide exchange; a mechanism for GTP-induced changes in effector and receptor binding surfaces; and a mechanism for GTPase activity not evident from previous data.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06510.
Organizational Affiliation: