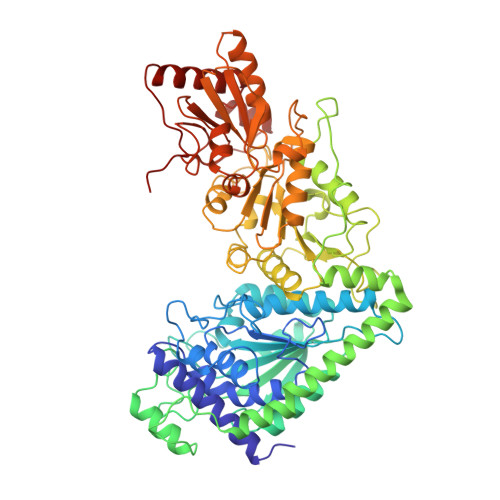
Specificity of coenzyme binding in thiamin diphosphate-dependent enzymes. Crystal structures of yeast transketolase in complex with analogs of thiamin diphosphate.
Konig, S., Schellenberger, A., Neef, H., Schneider, G.(1994) J Biological Chem 269: 10879-10882
- PubMed: 8144674 Search on PubMed
- Primary Citation Related Structures:
1TKA, 1TKB, 1TKC - PubMed Abstract:
The three-dimensional structures of complexes of yeast apotransketolase with the coenzyme analogs 6'-methyl, N1'-pyridyl, and N3'-pyridyl thiamin diphosphate, respectively, were determined with protein crystallographic methods. All three coenzyme analogs bind to the enzyme in a fashion highly similar to the cofactor thiamin diphosphate. Thus, either one of the hydrogen bonds of the pyrimidine ring nitrogens to the protein is sufficient for proper binding and positioning of the cofactor. The lack of catalytic activity of the N3'-pyridyl analog is not due to incorrect orientation of the pyrimidine ring, but results from the absence of the hydrogen bond between the N1' nitrogen atom and the conserved residue Glu418. The structure analysis provides further evidence for the importance of this conserved interaction for enzymatic thiamin catalysis.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala.
Organizational Affiliation: