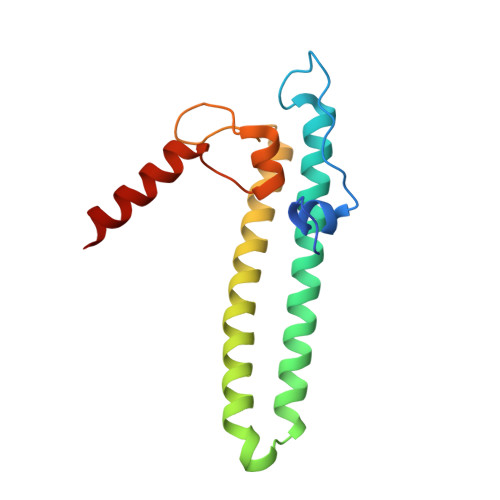
Regulation through the secondary channel--structural framework for ppGpp-DksA synergism during transcription
Perederina, A., Svetlov, V., Vassylyeva, M.N., Tahirov, T.H., Yokoyama, S., Artsimovitch, I., Vassylyev, D.G.(2004) Cell 118: 297-309
- PubMed: 15294156 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2004.06.030
- Primary Citation Related Structures:
1TJL - PubMed Abstract:
Bacterial transcription is regulated by the alarmone ppGpp, which binds near the catalytic site of RNA polymerase (RNAP) and modulates its activity. We show that the DksA protein is a crucial component of ppGpp-dependent regulation. The 2.0 A resolution structure of Escherichia coli DksA reveals a globular domain and a coiled coil with two highly conserved Asp residues at its tip that is reminiscent of the transcript cleavage factor GreA. This structural similarity suggests that DksA coiled coil protrudes into the RNAP secondary channel to coordinate a ppGpp bound Mg2+ ion with the Asp residues, thereby stabilizing the ppGpp-RNAP complex. Biochemical analysis demonstrates that DksA affects transcript elongation, albeit differently from GreA; augments ppGpp effects on initiation; and binds directly to RNAP, positioning the Asp residues near the active site. Substitution of these residues eliminates the synergy between DksA and ppGpp. Thus, the secondary channel emerges as a common regulatory entrance for transcription factors.
- Cellular Signaling Laboratory, Mikazuki-cho, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: