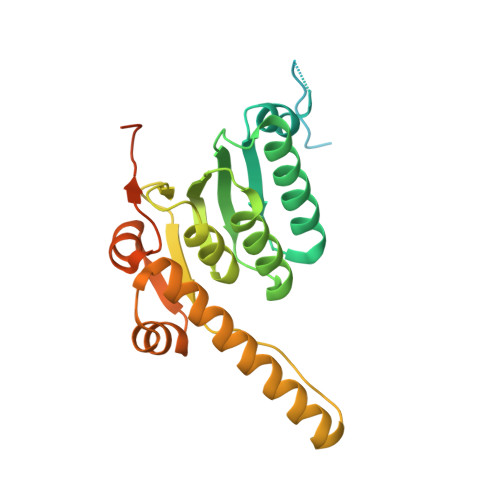
Crystallography and mutagenesis point to an essential role for the N-terminus of human mitochondrial ClpP
Kang, S.G., Maurizi, M.R., Thompson, M., Mueser, T., Ahvazi, B.(2004) J Struct Biol 148: 338-352
- PubMed: 15522782 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2004.07.004
- Primary Citation Related Structures:
1TG6 - PubMed Abstract:
We have determined a 2.1 A crystal structure for human mitochondrial ClpP (hClpP), the proteolytic component of the ATP-dependent ClpXP protease. HClpP has a structure similar to that of the bacterial enzyme, with the proteolytic active sites sequestered within an aqueous chamber formed by face-to-face assembly of the two heptameric rings. The hydrophobic N-terminal peptides of the subunits are bound within the narrow (12 A) axial channel, positioned to interact with unfolded substrates translocated there by the associated ClpX chaperone. Mutation or deletion of these residues causes a drastic decrease in ClpX-mediated protein and peptide degradation. Residues 8-16 form a mobile loop that extends above the ring surface and is also required for activity. The 28 amino acid C-terminal domain, a unique feature of mammalian ClpP proteins, lies on the periphery of the ring, with its proximal portion forming a loop that extends out from the ring surface. Residues at the start of the C-terminal domain impinge on subunit interfaces within the ring and affect heptamer assembly and stability. We propose that the N-terminal peptide of ClpP is a structural component of the substrate translocation channel and may play an important functional role as well.
- Laboratory of Cell Biology, National Cancer Institute, Bethesda, MD 20892-4255, USA.
Organizational Affiliation: