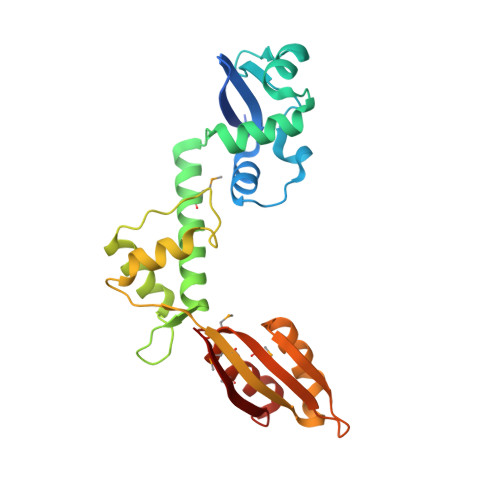
Structural and mutational analysis of the SBDS protein family. Insight into the leukemia-associated Shwachman-Diamond Syndrome.
Shammas, C., Menne, T.F., Hilcenko, C., Michell, S.R., Goyenechea, B., Boocock, G.R., Durie, P.R., Rommens, J.M., Warren, A.J.(2005) J Biological Chem 280: 19221-19229
- PubMed: 15701631 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M414656200
- Primary Citation Related Structures:
1T95 - PubMed Abstract:
Shwachman-Diamond Syndrome (SDS) is an autosomal recessive disorder characterized by bone marrow failure with significant predisposition to the development of poor prognosis myelodysplasia and leukemia, exocrine pancreatic failure and metaphyseal chondrodysplasia. Although the SBDS gene mutated in this disorder is highly conserved in Archaea and all eukaryotes, the function is unknown. To interpret the molecular consequences of SDS-associated mutations, we have solved the crystal structure of the Archaeoglobus fulgidus SBDS protein orthologue at a resolution of 1.9 angstroms, revealing a three domain architecture. The N-terminal (FYSH) domain is the most frequent target for disease mutations and contains a novel mixed alpha/beta-fold identical to the single domain yeast protein Yhr087wp that is implicated in RNA metabolism. The central domain consists of a three-helical bundle, whereas the C-terminal domain has a ferredoxin-like fold. By genetic complementation analysis of the essential Saccharomyces cerevisiae SBDS orthologue YLR022C, we demonstrate an essential role in vivo for the FYSH domain and the central three-helical bundle. We further show that the common SDS-related K62X truncation is non-functional. Most SDS-related missense mutations that alter surface epitopes do not impair YLR022C function, but mutations affecting residues buried in the hydrophobic core of the FYSH domain severely impair or abrogate complementation. These data are consistent with absence of homozygosity for the common K62X truncation mutation in individuals with SDS, indicating that the SDS disease phenotype is a consequence of expression of hypomorphic SBDS alleles and that complete loss of SBDS function is likely to be lethal.
- Medical Research Council Laboratory of Molecular Biology, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: