Crystal structure of ADP bound FAD synthetase
Shin, D.H., Wang, W., Kim, R., Yokota, H., Kim, S.-H.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| riboflavin kinase/FMN adenylyltransferase | 293 | Thermotoga maritima | Mutation(s): 0 EC: 2.7.7.2 (UniProt), 2.7.1.26 (UniProt) | 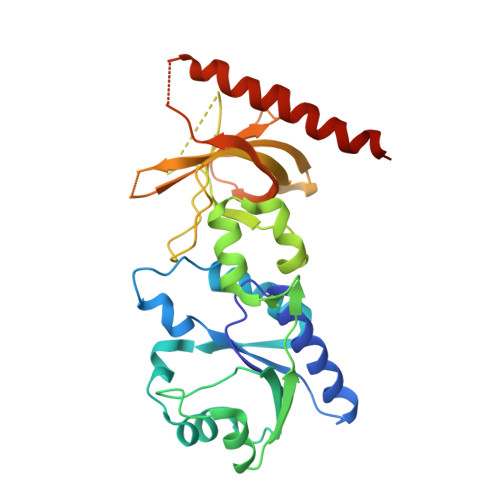 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9WZW1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| FMN Download:Ideal Coordinates CCD File | E [auth A], H [auth B] | FLAVIN MONONUCLEOTIDE C17 H21 N4 O9 P FVTCRASFADXXNN-SCRDCRAPSA-N |  | ||
| ADP Download:Ideal Coordinates CCD File | C [auth A], F [auth B] | ADENOSINE-5'-DIPHOSPHATE C10 H15 N5 O10 P2 XTWYTFMLZFPYCI-KQYNXXCUSA-N |  | ||
| AMP Download:Ideal Coordinates CCD File | D [auth A], G [auth B] | ADENOSINE MONOPHOSPHATE C10 H14 N5 O7 P UDMBCSSLTHHNCD-KQYNXXCUSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 67.803 | α = 90 |
| b = 82.661 | β = 116.42 |
| c = 66.717 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| CNS | refinement |
| CNS | phasing |