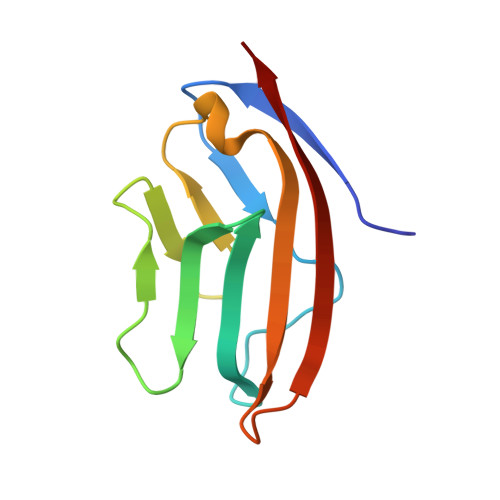
Design of a calcium-binding protein with desired structure in a cell adhesion molecule.
Yang, W., Wilkins, A.L., Ye, Y., Liu, Z.R., Li, S.Y., Urbauer, J.L., Hellinga, H.W., Kearney, A., van der Merwe, P.A., Yang, J.J.(2005) J Am Chem Soc 127: 2085-2093
- PubMed: 15713084 Search on PubMed
- DOI: https://doi.org/10.1021/ja0431307
- Primary Citation Related Structures:
1T6W - PubMed Abstract:
Ca2+, "a signal of life and death", controls numerous cellular processes through interactions with proteins. An effective approach to understanding the role of Ca2+ is the design of a Ca2+-binding protein with predicted structural and functional properties. To design de novo Ca2+-binding sites in proteins is challenging due to the high coordination numbers and the incorporation of charged ligand residues, in addition to Ca2+-induced conformational change. Here, we demonstrate the successful design of a Ca2+-binding site in the non-Ca2+-binding cell adhesion protein CD2. This designed protein, Ca.CD2, exhibits selectivity for Ca2+ versus other di- and monovalent cations. In addition, La3+ (Kd 5.0 microM) and Tb3+ (Kd 6.6 microM) bind to the designed protein somewhat more tightly than does Ca2+ (Kd 1.4 mM). More interestingly, Ca.CD2 retains the native ability to associate with the natural target molecule. The solution structure reveals that Ca.CD2 binds Ca2+ at the intended site with the designed arrangement, which validates our general strategy for designing de novo Ca2+-binding proteins. The structural information also provides a close view of structural determinants that are necessary for a functional protein to accommodate the metal-binding site. This first success in designing Ca2+-binding proteins with desired structural and functional properties opens a new avenue in unveiling key determinants to Ca2+ binding, the mechanism of Ca2+ signaling, and Ca2+-dependent cell adhesion, while avoiding the complexities of the global conformational changes and cooperativity in natural Ca2+-binding proteins. It also represents a major achievement toward designing functional proteins controlled by Ca2+ binding.
- Department of Chemistry, Center for Drug Design and Biotechnology, Georgia State University, Atlanta, Georgia 30303, USA.
Organizational Affiliation: