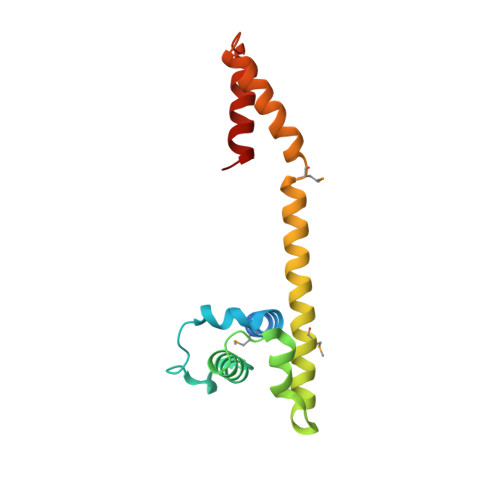
Crystal and solution structures of the helicase-binding domain of Escherichia coli primase
Oakley, A.J., Loscha, K.V., Schaeffer, P.M., Liepinsh, E., Pintacuda, G., Wilce, M.C.J., Otting, G., Dixon, N.E.(2005) J Biological Chem 280: 11495-11504
- PubMed: 15649896 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M412645200
- Primary Citation Related Structures:
1T3W - PubMed Abstract:
During bacterial DNA replication, the DnaG primase interacts with the hexameric DnaB helicase to synthesize RNA primers for extension by DNA polymerase. In Escherichia coli, this occurs by transient interaction of primase with the helicase. Here we demonstrate directly by surface plasmon resonance that the C-terminal domain of primase is responsible for interaction with DnaB6. Determination of the 2.8-angstroms crystal structure of the C-terminal domain of primase revealed an asymmetric dimer. The monomers have an N-terminal helix bundle similar to the N-terminal domain of DnaB, followed by a long helix that connects to a C-terminal helix hairpin. The connecting helix is interrupted differently in the two monomers. Solution studies using NMR showed that an equilibrium exists between a monomeric species with an intact, extended but naked, connecting helix and a dimer in which this helix is interrupted in the same way as in one of the crystal conformers. The other conformer is not significantly populated in solution, and its presence in the crystal is due largely to crystal packing forces. It is proposed that the connecting helix contributes necessary structural flexibility in the primase-helicase complex at replication forks.
- Research School of Chemistry, Australian National University, Canberra, Australian Capital Territory 0200, Australia.
Organizational Affiliation: