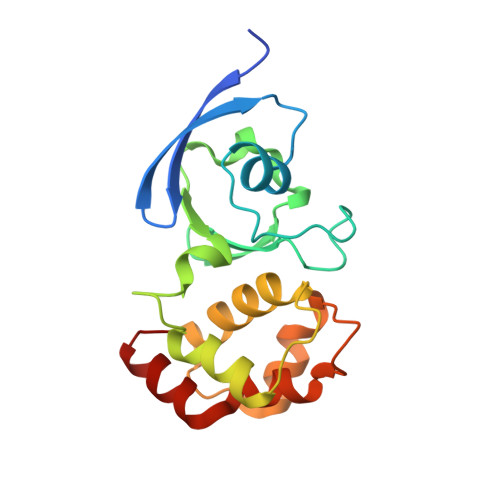
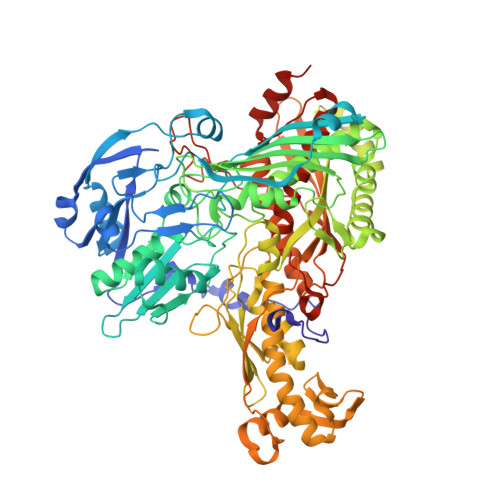
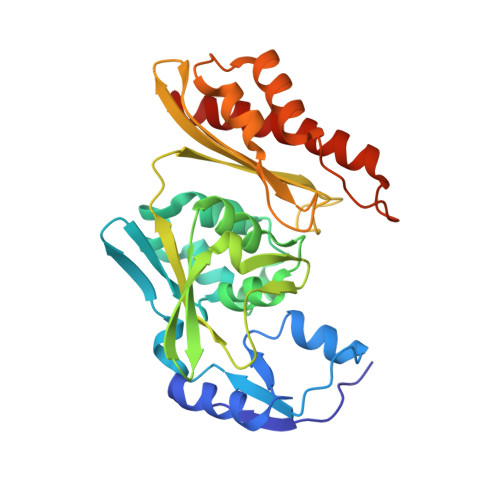
Active site geometry and substrate recognition of the molybdenum hydroxylase quinoline 2-oxidoreductase.
Bonin, I., Martins, B.M., Purvanov, V., Fetzner, S., Huber, R., Dobbek, H.(2004) Structure 12: 1425-1435
- PubMed: 15296736 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.05.014
- Primary Citation Related Structures:
1T3Q - PubMed Abstract:
The soil bacterium Pseudomonas putida 86 uses quinoline as a sole source of carbon and energy. Quinoline 2-oxidoreductase (Qor) catalyzes the first metabolic step converting quinoline to 2-oxo-1,2-dihydroquinoline. Qor is a member of the molybdenum hydroxylases. The molybdenum ion is coordinated by two ene-dithiolate sulfur atoms, two oxo-ligands, and a catalytically crucial sulfido-ligand, whose position in the active site was controversial. The 1.8 A resolution crystal structure of Qor indicates that the sulfido-ligand occupies the equatorial position at the molybdenum ion. The structural comparison of Qor with the allopurinol-inhibited xanthine dehydrogenase from Rhodobacter capsulatus allows direct insight into the mechanism of substrate recognition and the identification of putative catalytic residues. The active site protein variants QorE743V and QorE743D were analyzed to assess the catalytic role of E743.
- Abteilung für Strukturforschung, Max-Planck-Institut für Biochemie, 82152 Martinsried, Germany.
Organizational Affiliation: