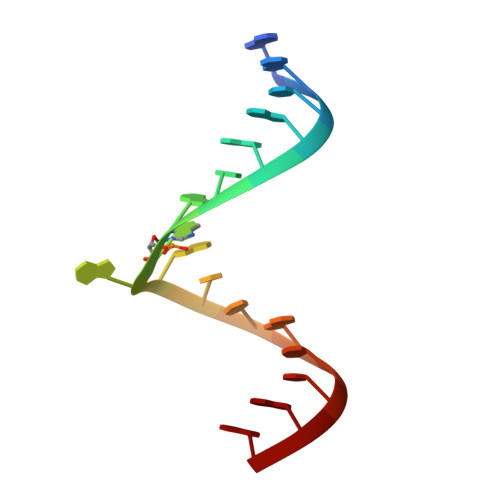
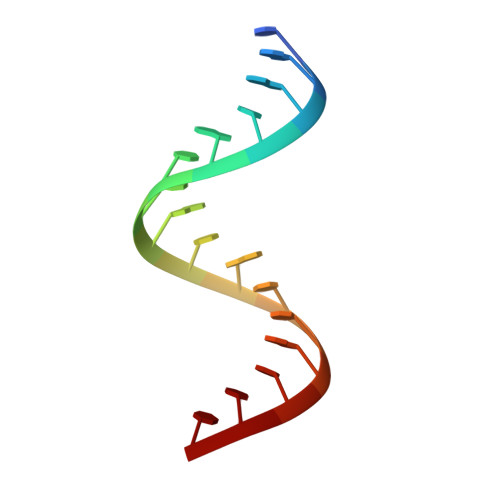
Monitoring molecular recognition of the ribosomal decoding site.
Shandrick, S., Zhao, Q., Han, Q., Ayida, B.K., Takahashi, M., Winters, G.C., Simonsen, K.B., Vourloumis, D., Hermann, T.(2004) Angew Chem Int Ed Engl 43: 3177-3182
- PubMed: 15199571 Search on PubMed
- DOI: https://doi.org/10.1002/anie.200454217
- Primary Citation Related Structures:
1T0D, 1T0E - Department of Structural Chemistry, Anadys Pharmaceuticals, Inc 9050 Camino Santa Fe, San Diego, CA 92121, USA.
Organizational Affiliation: