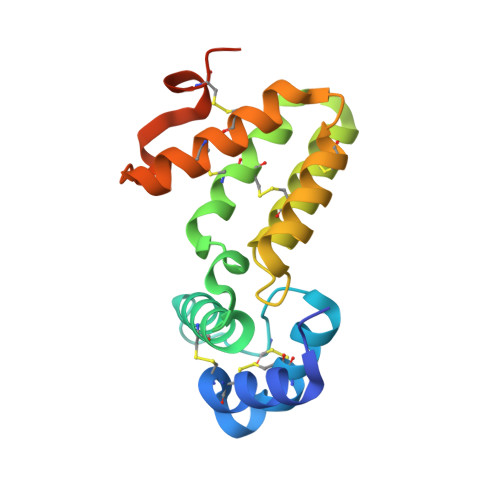
Crystal structure of Caenorhabditis elegans HER-1 and characterization of the interaction between HER-1 and TRA-2A.
Hamaoka, B.Y., Dann III, C.E., Geisbrecht, B.V., Leahy, D.J.(2004) Proc Natl Acad Sci U S A 101: 1673-11678
- PubMed: 15289613 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0402559101
- Primary Citation Related Structures:
1SZH - PubMed Abstract:
HER-1 is a secreted protein that promotes male development in the nematode Caenorhabditis elegans. HER-1 inhibits the function of TRA-2A, a multipass integral membrane protein thought to serve as its receptor. We report here the 1.5-A crystal structure of HER-1. The structure was solved by the multiwavelength anomalous diffraction method by using selenomethionyl-substituted HER-1 produced in Chinese hamster ovary cells. The HER-1 structure consists of two all-helical domains and is not closely homologous to any known structure. Sites of amino acid substitutions known to impair HER-1 function were mapped on the HER-1 structure and classified according to the likely mechanism by which they affect HER-1 activity. A subset of these and other amino acid substitutions on the HER-1 surface were assayed for their ability to disrupt interactions between HER-1 and TRA-2A-expressing cells, and a localized region on the HER-1 surface important for mediating this interaction was identified.
- Department of Biophysics and Biophysical Chemistry and Howard Hughes Medical Institute, The Johns Hopkins University School of Medicine, 725 North Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: