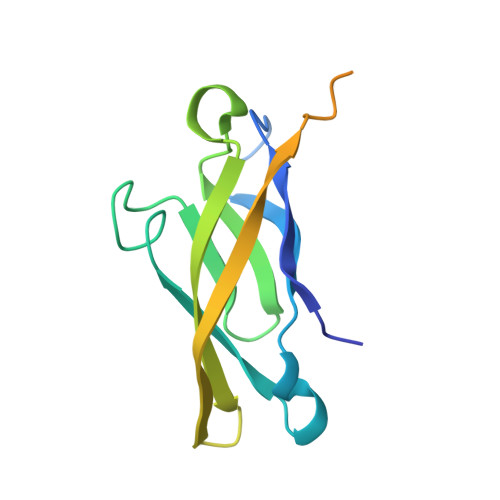
1H, 15N and 13C resonance assignments and monomeric structure of the amino-terminal extracellular domain of epithelial cadherin.
Overduin, M., Tong, K.I., Kay, C.M., Ikura, M.(1996) J Biomol NMR 7: 173-189
- PubMed: 8785495 Search on PubMed
- DOI: https://doi.org/10.1007/BF00202035
- Primary Citation Related Structures:
1SUH - PubMed Abstract:
E-cadherin is a transmembrane protein that provides Ca(2+)-dependent cell adhesion to epithelial cells. The large majority of the 1H, 15N, 13C and 13CO resonances of a 146-amino acid polypeptide from epithelial (E-) cadherin have been assigned using multidimensional NMR spectroscopy. The structure of the amino-terminal 100 amino acids, corresponding to the first extracellular repeat of E-cadherin [Overduin et al. (1995) Science, 267, 386-389], has been refined. The monomeric state of this isolated domain is demonstrated by light scattering and sedimentation analysis. Seven beta-strands and two short helices were identified by patterns of NOE cross-peaks, vicinal coupling constants and chemical shift indices. A novel structural motif termed a quasi-beta-helix found in the crystal structure of a neural (N-) cadherin domain [Shapiro et al. (1995) Nature, 374, 327-337] is characterized in detail for the first time by NMR. Slowly exchanging amides were concentrated in the beta-sheet region and quasi-beta-helix. The beta-barrel fold of the cadherin domain is topologically similar to the immunoglobulin fold. Comparison of this solution structure to the crystallized dimers of the N-terminal pair of E-cadherin domains [Nagar et al. (1996) Nature, 380, 360-364] and of the homologous single domain of N-cadherin reveals a conserved cadherin fold with minor structural differences, which can be accounted for by differences in metal ligation and oligomeric state.
- Division of Molecular and Structural Biology, Ontario Cancer Institute, Toronto, Canada.
Organizational Affiliation: