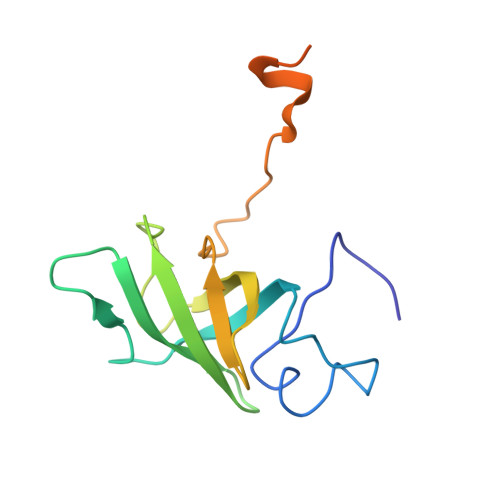
NMR Structure of the Heme Chaperone Ccme Reveals a Novel Functional Motif
Enggist, E., Thony-Meyer, L., Guntert, P., Pervushin, K.(2002) Structure 10: 1551-1557
- PubMed: 12429096 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00885-7
- Primary Citation Related Structures:
1SR3 - PubMed Abstract:
The concept of metal chaperones involves transient binding of metallic cofactors by specific proteins for delivery to enzymes in which they function. Metal chaperones thus provide a protective, as well as a transport, function. We report the first structure of a heme chaperone, CcmE, which comprises these two functions. We propose that the covalent attachment of heme to an exposed histidine occurs after heme binding at the surface of a rigid molecule with a flexible C-terminal domain. CcmE belongs to a family of proteins with a specific fold, which all share a function in delivery of specific molecular cargo.
- Institut für Mikrobiologie, Eidgenössische Technische Hochschule, Zürich, Switzerland.
Organizational Affiliation: