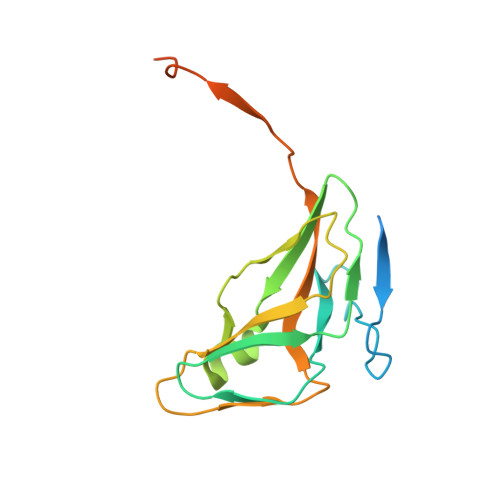
Crystal structure of the Mycobacterium tuberculosis dUTPase: insights into the catalytic mechanism.
Chan, S., Segelke, B., Lekin, T., Krupka, H., Cho, U.S., Kim, M.-Y., So, M., Kim, C.-Y., Naranjo, C.M., Rogers, Y.C., Park, M.S., Waldo, G.S., Pashkov, I., Cascio, D., Perry, J.L., Sawaya, M.R.(2004) J Mol Biology 341: 503-517
- PubMed: 15276840 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.06.028
- Primary Citation Related Structures:
1MQ7, 1SIX, 1SJN, 1SLH, 1SM8, 1SMC, 1SNF - PubMed Abstract:
The structure of Mycobacterium tuberculosis dUTP nucleotidohydrolase (dUTPase) has been determined at 1.3 Angstrom resolution in complex with magnesium ion and the non-hydrolyzable substrate analog, alpha,beta-imido dUTP. dUTPase is an enzyme essential for depleting potentially toxic concentrations of dUTP in the cell. Given the importance of its biological role, it has been proposed that inhibiting M.tuberculosis dUTPase might be an effective means to treat tuberculosis infection in humans. The crystal structure presented here offers some insight into the potential for designing a specific inhibitor of the M.tuberculosis dUTPase enzyme. The structure also offers new insights into the mechanism of dUTP hydrolysis by providing an accurate representation of the enzyme-substrate complex in which both the metal ion and dUTP analog are included. The structure suggests that inclusion of a magnesium ion is important for stabilizing the position of the alpha-phosphorus for an in-line nucleophilic attack. In the absence of magnesium, the alpha-phosphate of dUTP can have either of the two positions which differ by 4.5 Angstrom. A transiently ordered C-terminal loop further assists catalysis by shielding the general base, Asp83, from solvent thus elevating its pK(a) so that it might in turn activate a tightly bound water molecule for nucleophilic attack. The metal ion coordinates alpha, beta, and gamma phosphate groups with tridentate geometry identical with that observed in the crystal structure of DNA polymerase beta complexed with magnesium and dNTP analog, revealing some common features in catalytic mechanism.
- UCLA-DOE Laboratory of Structural Biology and Molecular Medicine, 206 Boyer Hall, Box 951570, Los Angeles, CA 90095-1570, USA.
Organizational Affiliation: