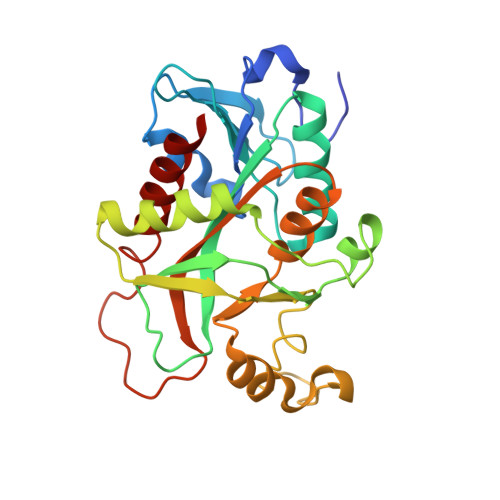
Preliminary investigation of the three-dimensional structure of Salmonella typhimurium uridine phosphorylase in the crystalline state.
Dontsova, M.V., Gabdoulkhakov, A.G., Molchan, O.K., Lashkov, A.A., Garber, M.B., Mironov, A.S., Zhukhlistova, N.E., Morgunova, E.Y., Voelter, W., Betzel, C., Zhang, Y., Ealick, S.E., Mikhailov, A.M.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 337-340
- PubMed: 16511035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309105007463
- Primary Citation Related Structures:
1SJ9 - PubMed Abstract:
Uridine phosphorylase (UPh) catalyzes the phosphorolytic cleavage of the C-N glycosidic bond of uridine to ribose 1-phosphate and uracil in the pyrimidine-salvage pathway. The crystal structure of the Salmonella typhimurium uridine phosphorylase (StUPh) has been determined at 2.5 A resolution and refined to an R factor of 22.1% and an Rfree of 27.9%. The hexameric StUPh displays 32 point-group symmetry and utilizes both twofold and threefold non-crystallographic axes. A phosphate is bound at the active site and forms hydrogen bonds to Arg91, Arg30, Thr94 and Gly26 of one monomer and Arg48 of an adjacent monomer. The hexameric StUPh model reveals a close structural relationship to Escherichia coli uridine phosphorylase (EcUPh).
- Institute of Protein Research, Russian Academy of Sciences, Pushchino, Russia.
Organizational Affiliation: