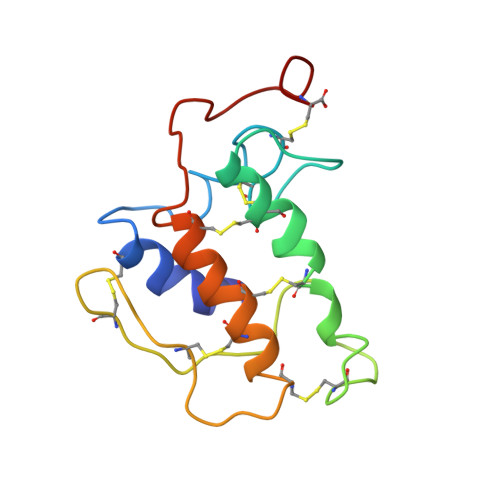
Solution structure of porcine pancreatic phospholipase A2 complexed with micelles and a competitive inhibitor.
van den Berg, B., Tessari, M., Boelens, R., Dijkman, R., Kaptein, R., de Haas, G.H., Verheij, H.M.(1995) J Biomol NMR 5: 110-121
- PubMed: 7703697 Search on PubMed
- DOI: https://doi.org/10.1007/BF00208802
- Primary Citation Related Structures:
1SFV, 1SFW - PubMed Abstract:
The three-dimensional structure of porcine pancreatic PLA2 (PLA2), present in a 40 kDa ternary complex with micelles and a competitive inhibitor, has been determined using multidimensional heteronuclear NMR spectroscopy. The structure of the protein (124 residues) is based on 1854 constraints, comprising 1792 distance and 62 phi torsion angle constraints. A total of 18 structures was calculated using a combined approach of distance geometry and restrained molecular dynamics. The atomic rms distribution about the mean coordinate positions for residues 1-62 and 72-124 is 0.75 +/- 0.09 A for the backbone atoms and 1.14 +/- 0.10 A for all atoms. The rms difference between the averaged minimized NMR structures of the free PLA2 and PLA2 in the ternary complex is 3.5 A for the backbone atoms and 4.0 A for all atoms. Large differences occur for the calcium-binding loop and the surface loop from residues 62 through 72. The most important difference is found for the first three residues of the N-terminal alpha-helix. Whereas free in solution Ala1, Leu2 and Trp3 are disordered, with the alpha-helical conformation with the alpha-amino group buried inside the protein. As a consequence, the important conserved hydrogen bonding network which is also seen in the crystal structures is present only in the ternary complex, but not in free PLA2. Thus, the NMR structure of the N-terminal region (as well as the calcium-binding loop and the surface loop) of PLA2 in the ternary complex resembles that of the crystal structure. Comparison of the NMR structures of the free enzyme and the enzyme in the ternary complex indicates that conformational changes play a role in the interfacial activation of PLA2.
- Center for Biomembranes and Lipid Enzymology, Utrecht University, The Netherlands.
Organizational Affiliation: