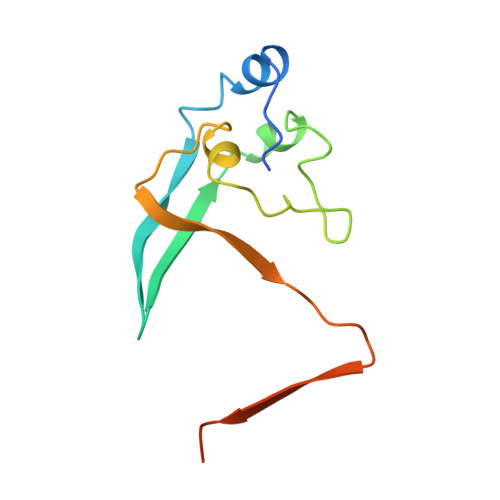
Crystal structure of the cell cycle-regulatory protein suc1 reveals a beta-hinge conformational switch.
Bourne, Y., Arvai, A.S., Bernstein, S.L., Watson, M.H., Reed, S.I., Endicott, J.E., Noble, M.E., Johnson, L.N., Tainer, J.A.(1995) Proc Natl Acad Sci U S A 92: 10232-10236
- PubMed: 7479758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.22.10232
- Primary Citation Related Structures:
1SCE - PubMed Abstract:
The Schizosaccharomyces pombe cell cycle-regulatory protein suc1, named as the suppressor of cdc2 temperature-sensitive mutations, is essential for cell cycle progression. To understand suc1 structure-function relationships and to help resolve conflicting interpretations of suc1 function based on genetic studies of suc1 and its functional homologs in both lower and higher eukaryotes, we have determined the crystal structure of the beta-interchanged suc1 dimer. Each domain consists of three alpha-helices and a four-stranded beta-sheet, completed by the interchange of terminal beta-strands between the two subunits. This beta-interchanged suc1 dimer, when compared with the beta-hairpin single-domain folds of suc1, reveals a beta-hinge motif formed by the conserved amino acid sequence HVPEPH. This beta-hinge mediates the subunit conformation and assembly of suc1: closing produces the intrasubunit beta-hairpin and single-domain fold, whereas opening leads to the intersubunit beta-strand interchange and interlocked dimer assembly reported here. This conformational switch markedly changes the surface accessibility of sequence-conserved residues available for recognition of cyclin-dependent kinase, suggesting a structural mechanism for beta-hinge-mediated regulation of suc1 biological function. Thus, suc1 belongs to the family of domain-swapping proteins, consisting of intertwined and dimeric protein structures in which the dual assembly modes regulate their function.
- Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: