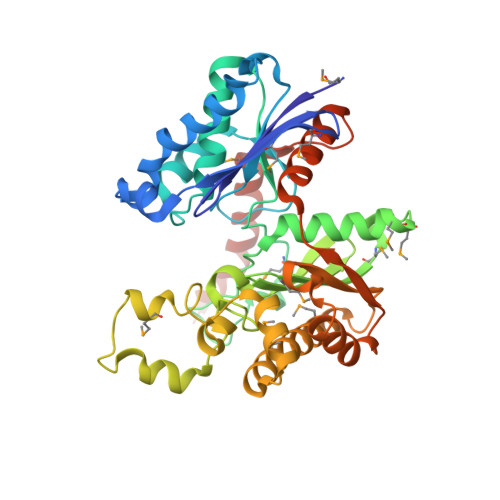
Crystal structure of butyrate kinase 2 from Thermotoga maritima, a member of the ASKHA superfamily of phosphotransferases.
Diao, J., Hasson, M.S.(2009) J Bacteriol 191: 2521-2529
- PubMed: 19201797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00906-08
- Primary Citation Related Structures:
1SAZ - PubMed Abstract:
The enzymatic transfer of phosphoryl groups is central to the control of many cellular processes. One of the phosphoryl transfer mechanisms, that of acetate kinase, is not completely understood. Besides better understanding of the mechanism of acetate kinase, knowledge of the structure of butyrate kinase 2 (Buk2) will aid in the interpretation of active-site structure and provide information on the structural basis of substrate specificity. The gene buk2 from Thermotoga maritima encodes a member of the ASKHA (acetate and sugar kinases/heat shock cognate/actin) superfamily of phosphotransferases. The encoded protein Buk2 catalyzes the phosphorylation of butyrate and isobutyrate. We have determined the 2.5-A crystal structure of Buk2 complexed with (beta,gamma-methylene) adenosine 5'-triphosphate. Buk2 folds like an open-shelled clam, with each of the two domains representing one of the two shells. In the open active-site cleft between the N- and C-terminal domains, the active-site residues consist of two histidines, two arginines, and a cluster of hydrophobic residues. The ATP binding region of Buk2 in the C-terminal domain consists of abundant glycines for nucleotide binding, and the ATP binding motif is similar to those of other members of the ASKHA superfamily. The enzyme exists as an octamer, in which four disulfide bonds form between intermolecular cysteines. Sequence alignment and structure superposition identify the simplicity of the monomeric Buk2 structure, a probable substrate binding site, the key residues in catalyzing phosphoryl transfer, and the substrate specificity differences among Buk2, acetate, and propionate kinases. The possible enzyme mechanisms are discussed.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA. diaojs@netscape.net
Organizational Affiliation: