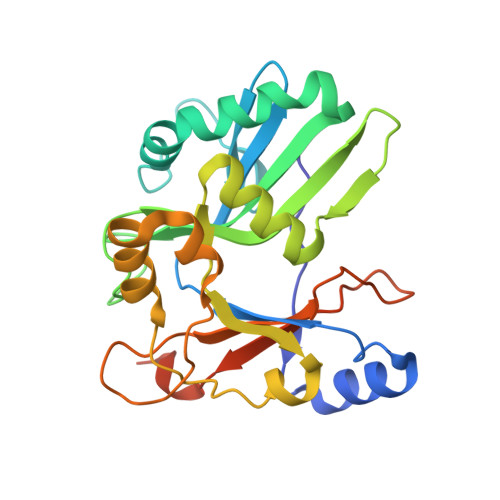
Structure and mechanism of RNA ligase.
Ho, C.K., Wang, L.K., Lima, C.D., Shuman, S.(2004) Structure 12: 327-339
- PubMed: 14962393 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.01.011
- Primary Citation Related Structures:
1S68 - PubMed Abstract:
T4 RNA ligase 2 (Rnl2) exemplifies an RNA ligase family that includes the RNA editing ligases (RELs) of Trypanosoma and Leishmania. The Rnl2/REL enzymes are defined by essential signature residues and a unique C-terminal domain, which we show is essential for sealing of 3'-OH and 5'-PO4 RNA ends by Rnl2, but not for ligase adenylation or phosphodiester bond formation at a preadenylated AppRNA end. The N-terminal segment Rnl2(1-249) of the 334 aa Rnl2 protein comprises an autonomous adenylyltransferase/AppRNA ligase domain. We report the 1.9 A crystal structure of the ligase domain with AMP bound at the active site, which reveals a shared fold, catalytic mechanism, and evolutionary history for RNA ligases, DNA ligases, and mRNA capping enzymes.
- Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10021, USA.
Organizational Affiliation: