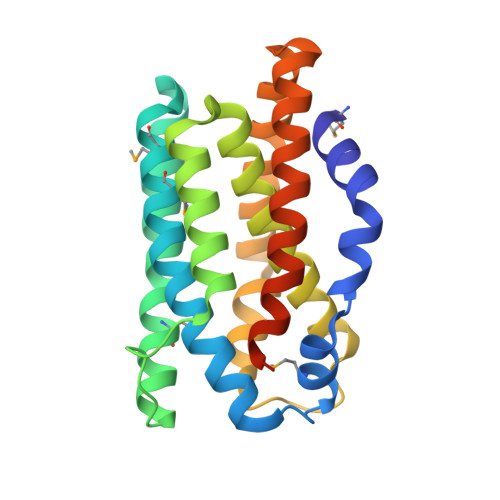
The 2.35 A structure of the TenA homolog from Pyrococcus furiosus supports an enzymatic function in thiamine metabolism.
Benach, J., Edstrom, W.C., Lee, I., Das, K., Cooper, B., Xiao, R., Liu, J., Rost, B., Acton, T.B., Montelione, G.T., Hunt, J.F.(2005) Acta Crystallogr D Biol Crystallogr 61: 589-598
- PubMed: 15858269 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905005147
- Primary Citation Related Structures:
1RTW - PubMed Abstract:
TenA (transcriptional enhancer A) has been proposed to function as a transcriptional regulator based on observed changes in gene-expression patterns when overexpressed in Bacillus subtilis. However, studies of the distribution of proteins involved in thiamine biosynthesis in different fully sequenced genomes have suggested that TenA may be an enzyme involved in thiamine biosynthesis, with a function related to that of the ThiC protein. The crystal structure of PF1337, the TenA homolog from Pyrococcus furiosus, is presented here. The protomer comprises a bundle of alpha-helices with a similar tertiary structure and topology to that of human heme oxygenase-1, even though there is no significant sequence homology. A solvent-sequestered cavity lined by phylogenetically conserved residues is found at the core of this bundle in PF1337 and this cavity is observed to contain electron density for 4-amino-5-hydroxymethyl-2-methylpyrimidine phosphate, the product of the ThiC enzyme. In contrast, the modestly acidic surface of PF1337 shows minimal levels of sequence conservation and a dearth of the basic residues that are typically involved in DNA binding in transcription factors. Without significant conservation of its surface properties, TenA is unlikely to mediate functionally important protein-protein or protein-DNA interactions. Therefore, the crystal structure of PF1337 supports the hypothesis that TenA homologs have an indirect effect in altering gene-expression patterns and function instead as enzymes involved in thiamine metabolism.
- Department of Biological Sciences and Northeast Structural Genomics Consortium, 702A Fairchild Center, MC2434, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: