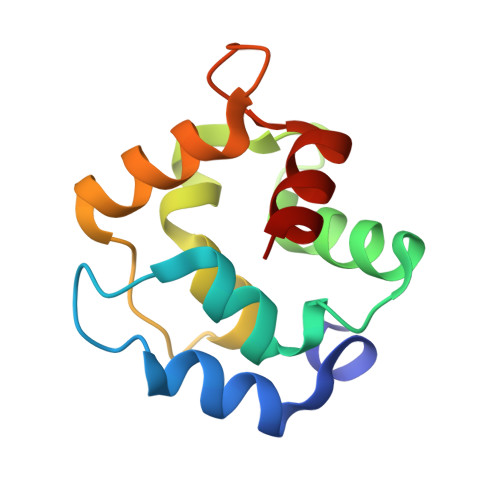
Refined crystal structure of rat parvalbumin, a mammalian alpha-lineage parvalbumin, at 2.0 A resolution.
McPhalen, C.A., Sielecki, A.R., Santarsiero, B.D., James, M.N.(1994) J Mol Biology 235: 718-732
- PubMed: 8289291 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1023
- Primary Citation Related Structures:
1RTP - PubMed Abstract:
We present here the X-ray crystal structure of the rat alpha-parvalbumin from fast twitch muscle. This protein (M(r) 11.8 kDa) crystallizes in space group P2(1)2(1)2(1) with unit cell dimensions of a = 34.3 A, b = 55.0 A, c = 156.1 A and three molecules in the asymmetric unit. The protein structure was solved by the molecular replacement method and has been refined to a crystallographic R-factor [formula: see text] of 0.181 for all reflections with I/sigma(I) > or = 2 (I = intensity) between 8.0 and 2.0 A resolution. The molecules located most easily in the molecular replacement rotation function had lower overall thermal motion parameters and higher numbers of intermolecular crystal packing contacts. The overall fold of the polypeptide chain for the rat alpha-parvalbumin is similar to other known parvalbumin structures (root-mean-square deviations in alpha-carbon atom positions range from 0.60 to 0.87 A). There are two Ca(2+)-binding sites in parvalbumins, and there is some evidence for a third ion-binding site, adjacent to the CD site, in the rat species. The level of structural variability among the best-ordered regions of the three independent rat alpha-parvalbumin molecules in the crystallographic asymmetric unit is two to three times higher than the mean coordinate error (0.10 A), indicating flexibility in the molecule. Sequence differences between alpha and beta-lineage parvalbumins result in repacking of the hydrophobic core and some shifts in the protein backbone. The shifts are localized, however, and entire helices do not shift as rigid units.
- Department of Biochemistry, University of Alberta, Edmonton, Canada.
Organizational Affiliation: