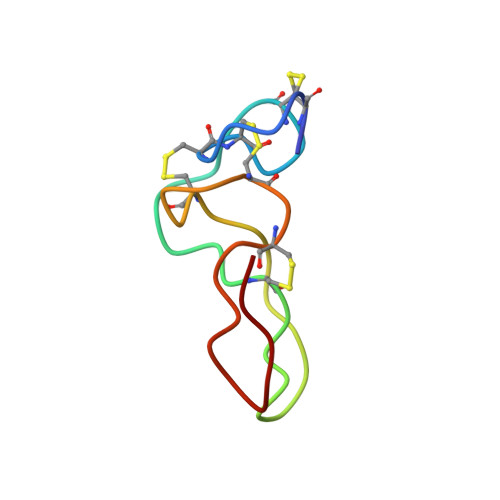
Conformation and concerted dynamics of the integrin-binding site and the C-terminal region of echistatin revealed by homonuclear NMR
Monleon, D., Esteve, V., Kovacs, H., Calvete, J.J., Celda, B.(2005) Biochem J 387: 57-66
- PubMed: 15535803 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20041343
- Primary Citation Related Structures:
1RO3 - PubMed Abstract:
Echistatin is a potent antagonist of the integrins alpha(v)beta3, alpha5beta1 and alpha(IIb)beta3. Its full inhibitory activity depends on an RGD (Arg-Gly-Asp) motif expressed at the tip of the integrin-binding loop and on its C-terminal tail. Previous NMR structures of echistatin showed a poorly defined integrin-recognition sequence and an incomplete C-terminal tail, which left the molecular basis of the functional synergy between the RGD loop and the C-terminal region unresolved. We report a high-resolution structure of echistatin and an analysis of its internal motions by off-resonance ROESY (rotating-frame Overhauser enhancement spectroscopy). The full-length C-terminal polypeptide is visible as a beta-hairpin running parallel to the RGD loop and exposing at the tip residues Pro43, His44 and Lys45. The side chains of the amino acids of the RGD motif have well-defined conformations. The integrin-binding loop displays an overall movement with maximal amplitude of 30 degrees . Internal angular motions in the 100-300 ps timescale indicate increased flexibility for the backbone atoms at the base of the integrin-recognition loop. In addition, backbone atoms of the amino acids Ala23 (flanking the R24GD26 tripeptide) and Asp26 of the integrin-binding motif showed increased angular mobility, suggesting the existence of major and minor hinge effects at the base and the tip, respectively, of the RGD loop. A strong network of NOEs (nuclear Overhauser effects) between residues of the RGD loop and the C-terminal tail indicate concerted motions between these two functional regions. A full-length echistatin-alpha(v)beta3 docking model suggests that echistatin's C-terminal amino acids may contact alpha(v)-subunit residues and provides new insights to delineate structure-function correlations.
- Departamento de Química Física, Universitat de València, Dr. Moliner 50, 46100 Burjassot, Valencia, Spain.
Organizational Affiliation: