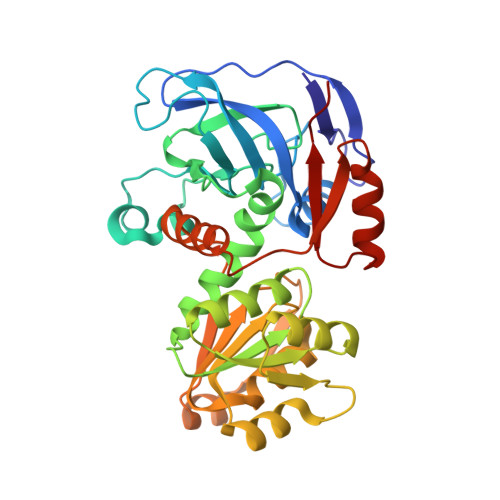
Crystal Structure and Amide H/D Exchange of Binary Complexes of Alcohol Dehydrogenase from Bacillus stearothermophilus: Insight into Thermostability and Cofactor Binding
Ceccarelli, C., Liang, Z.X., Strickler, M., Prehna, G., Goldstein, B.M., Klinman, J.P., Bahnson, B.J.(2004) Biochemistry 43: 5266-5277
- PubMed: 15122892 Search on PubMed
- DOI: https://doi.org/10.1021/bi049736p
- Primary Citation Related Structures:
1RJW - PubMed Abstract:
The crystal structure of NAD(+)-dependent alcohol dehydrogenase from Bacillus stearothermophilus strain LLD-R (htADH) was determined using X-ray diffraction data at a resolution of 2.35 A. The structure of homotetrameric htADH is highly homologous to those of bacterial and archaeal homotetrameric alcohol dehydrogenases (ADHs) and also to the mammalian dimeric ADHs. There is one catalytic zinc atom and one structural zinc atom per enzyme subunit. The enzyme was crystallized as a binary complex lacking the nicotinamide adenine dinucleotide (NAD(+)) cofactor but including a zinc-coordinated substrate analogue trifluoroethanol. The binary complex structure is in an open conformation similar to ADH structures without the bound cofactor. Features important for the thermostability of htADH are suggested by a comparison with a homologous mesophilic enzyme (55% identity), NAD(+)-dependent alcohol dehydrogenase from Escherichia coli. To gain insight into the conformational change triggered by NAD(+) binding, amide hydrogen-deuterium exchange of htADH, in the presence and absence of NAD(+), was studied by HPLC-coupled electrospray mass spectrometry. When the deuteron incorporation of the protein-derived peptides was analyzed, it was found that 9 of 21 peptides show some decrease in the level of deuteron incorporation upon NAD(+) binding, and another 4 peptides display slower exchange rates. With one exception (peptide number 8), none of the peptides that are altered by bound NAD(+) are in contact with the alcohol-substrate-binding pocket. Furthermore, peptides 5 and 8, which are located outside the NAD(+)-binding pocket, are notable by displaying changes upon NAD(+) binding. This suggests that the transition from the open to the closed conformation caused by cofactor binding has some long-range effects on the protein structure and dynamics.
- Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware 19716, USA.
Organizational Affiliation: