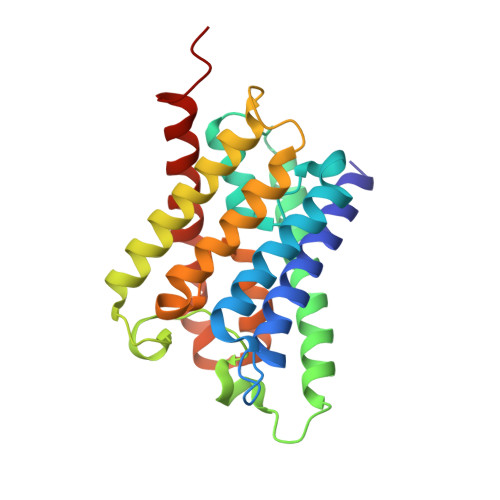
Architecture and selectivity in aquaporins: 2.5 a X-ray structure of aquaporin Z
Savage, D.F., Egea, P.F., Robles-Colmenares, Y., O'Connell III, J.D., Stroud, R.M.(2003) PLoS Biol 1: 334-340
- PubMed: 14691544 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.0000072
- Primary Citation Related Structures:
1RC2 - PubMed Abstract:
Aquaporins are a family of water and small molecule channels found in organisms ranging from bacteria to animals. One of these channels, the E. coli protein aquaporin Z (AqpZ), has been shown to selectively conduct only water at high rates. We have expressed, purified, crystallized, and solved the X-ray structure of AqpZ. The 2.5 A resolution structure of AqpZ suggests aquaporin selectivity results both from a steric mechanism due to pore size and from specific amino acid substitutions that regulate the preference for a hydrophobic or hydrophilic substrate. This structure provides direct evidence on the molecular mechanisms of specificity between water and glycerol in this family of channels from a single species. It is to our knowledge the first atomic resolution structure of a recombinant aquaporin and so provides a platform for combined genetic, mutational, functional, and structural determinations of the mechanisms of aquaporins and, more generally, the assembly of multimeric membrane proteins.
- Department of Biochemistry and Biophysics, University of California School of Medicine, San Francisco, California, USA.
Organizational Affiliation: