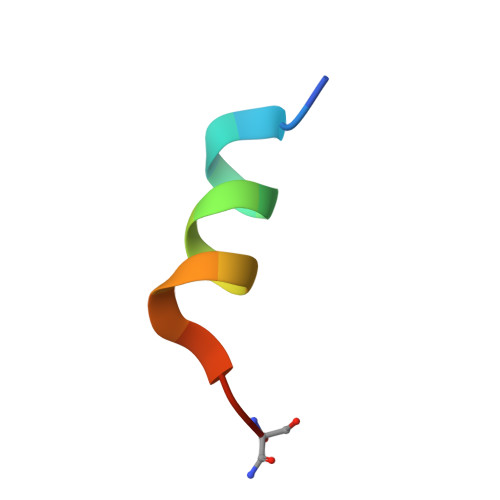
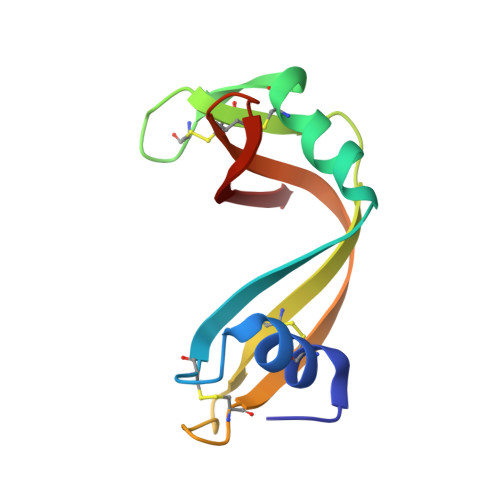
Crystallographic structures of ribonuclease S variants with nonpolar substitution at position 13: packing and cavities.
Varadarajan, R., Richards, F.M.(1992) Biochemistry 31: 12315-12327
- PubMed: 1463720 Search on PubMed
- DOI: https://doi.org/10.1021/bi00164a005
- Primary Citation Related Structures:
1RBC, 1RBD, 1RBE, 1RBF, 1RBG, 1RBH, 1RBI - PubMed Abstract:
Seven hydrophobic residues ranging in size from glycine to phenylalanine have been substituted for the wild-type methionine residue at position 13 in a 15-residue truncated version (S15) of S-peptide, the small component of ribonuclease S. Complexes of both S-15 and the seven variants with S-protein yielded isomorphous crystals. The structures of all eight complexes have been refined to final R-factors in the range of 17-19%. [See Kim, E. E. Varadarajan, R., Wyckoff, H. W., and Richards, F. M. (1992) Biochemistry (preceding paper in this issue) for the description of the reference S-15 complex.] Multiple side-chain conformations were seen for six residues in all of the complexes and for two to three additional residues in at least some of the complexes. Three of the complexes, Gly, Ala, and alpha-amino-n-butyric acid (ANB), contained a single water molecule in the cavity near residue 13 that makes three hydrogen bonds to protein atoms. Although space is available, no evidence for additional water in this region, ordered or disordered, was found. The atoms in the cavity wall tend to shrink the cavity by moving in on the small residues and to swell the cavity by moving out for the larger Phe substitution. A swelling seen with leucine was attributed to a shape effect since Leu, Ile, and Met all have the same volume. A slight volume contraction of the collection of interior residues outside of the region of position 13 was also noted. (All changes noted are in the direction to maintain a constant packing density averaged over the whole protein.) Leu51, a surface hydrophobic residue, moved considerably in the G, A, and ANB complexes in directionswhich would tend to decrease the cavity volume. The only other major change in position, 1.5 A, was the 66-69 loop, which is about 25 A from position 13. His12, Phe120, and Asp121 appear to be involved in this movement, but the connection with position 13 is not clear at all. The thermodynamic data on the association reaction for all of these complexes have been previously reported [Connelly, P. R., Varadarajan, R., Sturtevant, J. M., & Richards, F. M. (1990) Biochemistry 29, 6108-6114; Varadarajan, R., Connelly, P. R., Sturtevant, J. M., & Richards, F. M. (1992) Biochemistry 31, 1421-1426]. Some comments are offered on our initial attempts to correlate the structural changes with the changes in the thermodynamic parameters.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06511.
Organizational Affiliation: